Chemists craft molecule that self-assembles into flower-shaped crystalline patterns

The National Science Foundation has awarded $1.2 million to three research groups at Indiana University to advance research on self-assembling molecules and computer-aided design software required to create the next generation of solar cells, circuits, sensors and other technology.
This interdisciplinary team in the IU Bloomington College of Arts and Sciences' Department of Chemistry is led by Amar Flood, Steven Tait and Peter Ortoleva in collaboration with Mu-Hyun Baik of the Korea Advanced Institute of Science and Technology, who previously served at IU.
Designing new materials at the molecular level is a key goal of the U.S. government's Materials Genome Initiative, a project launched in 2011 to reduce the cost, and speed the creation, of these materials. As recipients of funds from the NSF's Designing Materials to Revolutionize and Engineer our Future program, the IU scientists will contribute to this national initiative.
"There are more than 100 million known molecules, but in the vast majority of cases we cannot predict what sort of structure they will form when those molecules start packing together," said Amar Flood, James F. Jackson Professor of Chemistry and Luther Dana Waterman Professor in the IU Bloomington Department of Chemistry, who is the principal investigator on the grant. "We want to be able to predict, as well as design, those structures."
The results would represent a "transformative approach to the discovery of organic materials," he said, combining computer-aided design, chemical synthesis and molecular characterization methods.
And recently, Flood and colleagues have shown such an ambitious goal is achievable.
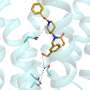
In a paper published Nov. 23 in Chemistry—A European Journal, the IU scientists describe an innovative "one-pot" method to synthesize a new macromolecule called a tricarbazolo triazolophane, or "tricarb."
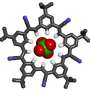
A multifunctional, ring-shaped structure, tricarb molecules bear alignment markers so that they line up perfectly with each other upon contact to form highly organized, multilayered patterns. Tricarb molecules also have a central pocket to capture the negatively charged particles known as anions.
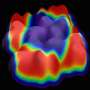
"Amar has developed a very elegant synthesis," said Steven Tait, an associate professor of chemistry who is a co-author on the paper and also a co-investigator on the NSF grant. "The result is molecules that recognize each other in a very specific way to order and stack in beautiful, flower-shaped crystalline patterns with potentially transformative properties."
The NSF-funded project will support creating molecular structures, like the tricarb molecule, that are specifically pre-programmed to self-assemble into three-dimensional structures that go beyond the comparatively simple, two-dimensional molecular arrangements.
"Creating building blocks that self-assemble into functional materials will be a major breakthrough in materials science and is a key component of the Materials Genome Initiative," said Stephen C. Jacobson, chair of the IU Bloomington Department of Chemistry. "I am pleased that the NSF has recognized our faculty's combined expertise in synthesis, characterization and theory in selecting them to contribute to this important initiative."
Specifically, Flood said, the ability to alternate different molecules in highly ordered patterns is a key step in creating organic electronics, a new class of material whose applications include highly efficient solar panels and advanced computer circuitry.
"The best solar cells right now are made of extremely pure silicon, which requires a very precise—and expensive—production process," Tait said. "But if we can create extremely pure, self-assembling organic materials, controlling the order of their interfaces and components at the molecular level, the performance of these organic materials will improve significantly, and their costs will go down."
Most important for the creation of new molecular structures, the IU team will use the grant to develop computer-aided design software enabling virtual experimentation with the potential to examine the millions of molecular compounds of interest to material scientists.
Currently, Flood explained, scientists must engage in an arduous and time-consuming process of trial-and-error to design new structures with highly specialized properties since no blueprint exists for how molecules will react upon coming into contact with each other. But with virtual experimentation, molecular engineers could screen 100 potential molecular combinations over 100 days, only then devoting time and resources to synthesize the top five candidates, which can itself require about 100 days per compound, creating enormous time-savings.
"CAD software is prevalent in electrical, mechanical and civil engineering, and we need that same technology at our fingertips for molecular and materials engineering," Flood said. "The innovations coming from our computational collaborators are key."
These collaborations are with Ortoleva, a Distinguished Professor in the IU Bloomington Department of Chemistry who will help develop the CAD software using recent advances in multi-scale simulation that employ Baik's work on atomic-level force fields.
"Ultimately, we plan to show experimentally how molecules can be programmed so that they assemble themselves into 2-D and 3-D arrangements, as well as produce a working, operational and accurate simulation software," Flood added. "Our goal will be to achieve high fidelity between theory, design and experimentation."
More information: Chemistry—A European Journal, onlinelibrary.wiley.com/doi/10 … /chem.201503161/full
Journal information: Chemistry – A European Journal
Provided by Indiana University