DNA sequencing shows divergent genomes in malaria vectors of Brazilian rainforest
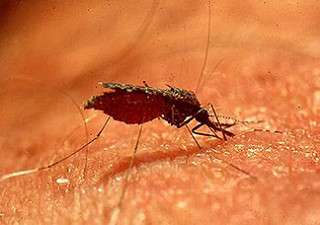
The Amazon rainforest occupies more than 2 million square miles (5.5 million square kilometers) in South America, 60% of it in Brazil. Far from being homogeneous, this vast region contains rivers and mountain ranges that foster biodiversity. Scientists are currently studying whether these natural barriers affect genomic diversity in Anopheles (Nyssorhynchus) darlingi Root, the primary malaria vector in this area.
A new study by Kevin J. Emerson, PhD, assistant professor of biology at St. Mary's College of Maryland and his international group of collaborators assessed the extent to which geographical barriers affected genetic variation among Anopheles darlingi populations. Such barriers may greatly influence the approaches used by scientists and physicians to control the spread of malaria throughout Brazil.
Until recently, research on malaria and the vectors for its transmission primarily targeted sub-Saharan Africa: For example, the genome of primary African vector, Anopheles gambiae, was sequenced in 2001, while that of Anopheles darlingi (a species in the Neotropics or "New World tropics" – mainly South and Central America) was finally sequenced more than a decade later, in 2014. Over the past decade, advances in DNA sequencing technologies have enabled genetics researchers to expand their focus from a few so-called "model organisms" to many organisms of interest.
"I think the focus for malaria vector research is starting to become more general, rather than focusing solely on one species of vector in Africa," said Dr. Emerson. "That's one of the reasons I like working on South American mosquitoes. With all of the focus on the genetics of African mosquitoes, the genetics involved in malaria transmission in the Neotropics has been relatively ignored. Research there is really just starting. And if we, as scientists, want to get to the basis of disease transmission, we must study diverse types of mosquitoes that can transmit malaria, rather than just one."
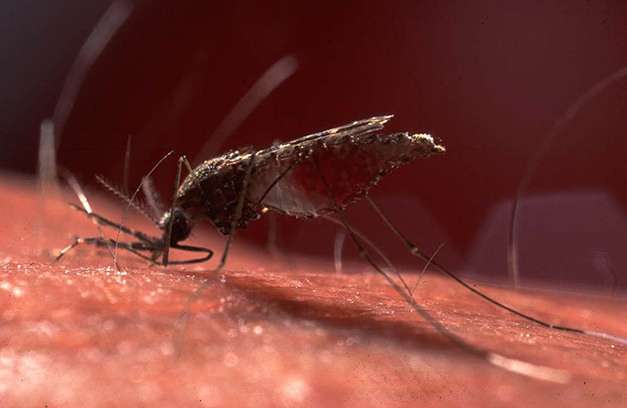
Drawing on his experience in evolutionary genetics, Dr. Emerson (with colleagues in Albany, NY, Eugene, OR, and São Paulo, Brazil) analyzed genomic DNA from 57 mosquitoes – pupae/larvae or adults collected from multiple habitats in each region to lower the risk of obtaining related individuals. Working at remote field sites in Amazonian Brazil, collectors placed specimens in tubes with desiccant to dry them as quickly as possible. DNA is stable in the tubes for up to a couple weeks, enough time to return to the university, preserve the specimens in alcohol, and eventually isolate their DNA.
The DNA content of these mosquitoes was then determined at 11,533 positions in their genomes, using a new genotyping technique (NextRAD genotyping) that can determine the specific DNA sequence for each individual at each position. Analysis of the genetic diversity among these individuals showed there were three distinct genetic clusters of indivdiuals (populations) that were geograpjhically isolated – mosquitoes in the Atlantic Forest province (Southeast), the Parana Forest province (West Atlantic forest), and the Brazilian dominion (Amazonian). After weighing their own evidence, and that of others, that the divergent genetic clusters may be well-separated species, the authors proposed that the West Atlantic cluster represents Anopheles paulistensis. This possibility would support species-level differentiation between the Atlantic Forest (Southeast) and Parana Forest (West Atlantic) clusters.
Phenotypic differences between these populations are too minor to be noticed by a classical taxonomist. "There has been some small discussion, based on wing shape, that these might be different species, but the evidence hasn't been very strong," said Dr. Emerson. But genome sequencing provides good supporting evidence for divergence. The same is true in Africa: "In Anopheles gambiae ... the species are identified genetically. You can't identify the different forms without knowing the genes. So that may be happening with the vector in the Neotropics as well."
The genetic divergence of these three Brazilian clusters pobably reflects ecological selection pressures as well as historical biogeographical processes that influenced contact and separation of these populations.
But beyond their immediate observations and proposals regarding mosquito speciation, the findings of Emerson and his colleagues may contribute significantly to disease control:
"When you're interested in disease transmission among humans, most of the research effort is put into understanding the disease as it exists in humans," said Dr. Emerson. "But one of the primary ways that the spread of these diseases has ever been controlled is by controlling the vectors of the disease, rather than controlling the disease after humans become infected." Although malaria is a human disease, it is caused by a parasite (of the genus Plasmodium) that is transmitted among humans by mosquitoes. Understanding the biology of the vector mosquito, and its interaction with the malaria parasite, may lead to control strategies.
For example mosquitoes in an area with very little malaria transmission may be quite genetically distinct from mosquitoes in a high malaria area. In terms of control, "part of the novelty in what we do is that we focus on the genetics and biology of the mosquito vector that transmits malaria, in order to identify those populations of mosquitoes causing the transmission," Emerson observed.
"If you think about malaria, we spend a lot of money and effort every year looking for drugs or vaccines, and they work a little bit. But the best way to prevent malaria transmission is sleeping under a bednet and reducing the occurrence of standing water that mosquitoes use to lay their eggs. That just shows how understanding the biology of the vector itself is critical in controlling disease transmission," noted Dr. Emerson. "The most effective disease controls have been at the level of the vector, and part of that is knowing what the vector is."
More information: Kevin J. Emerson et al. Brazilian Anopheles darlingi Root (Diptera: Culicidae) Clusters by Major Biogeographical Region, PLOS ONE (2015). DOI: 10.1371/journal.pone.0130773
Journal information: PLoS ONE
Provided by St. Mary's College of Maryland


















