The PLOS Comp Biol Macromolecular Structure and Dynamics collection
Living systems are in a state of perpetual motion. Down at the microscopic level, the atoms that make up biological macromolecules are, in the words of Richard Feynman, jiggling and wiggling. These atomic motions are beautifully choreographed and harnessed into productive events in the cell. Fluctuations of macromolecules between three-dimensional structures underlie all processes that maintain and replicate a living cell. Investigation of macromolecular structure and dynamics is fundamental to understanding how macromolecules carry out their functions in the cell.
Today, PLOS Computational Biology launches the new Macromolecular Structure and Dynamics Collection led by Ruth Nussinov (Editor-in-Chief) and Amarda Shehu (George Mason University, USA). Publishing in PLOS Computational Biology's 10th Anniversary year, the collection focuses on recent computational methods investigating various aspects of the relationship between macromolecular structure, dynamics, and function.
The Nobel Prize in Chemistry 2013, awarded to PLOS Computational Biology Michael Levitt (Associate Editor), Martin Karplus and Arieh Warshel "for the development of multiscale models for complex chemical systems", highlighted the central role that computation now plays in complementing wet-laboratory investigations of macromolecular structure, dynamics, and function. This collection aims to highlight some of the most recent advancements in computational treatments of macromolecules.
The focus is on novel methods capable of addressing the disparate spatial and time scales accessed by dynamic macromolecules and leading to new understanding and biological discoveries. Articles address a variety of methods such as multi-scale estimation of binding kinetics, mapping of the protein folding universe, quantitative comparison of conformational ensembles, and many more. For more details of these methods please see an outline in our introductory Editorial.
Through this collection we hope to convey and celebrate the tremendous progress that computational structural biology has made over the last two decades. Along with new, higher-resolution, and increasingly efficient high-throughput experimental techniques that are leading to huge amounts of accumulating data, computational biology is pushing the frontiers to meet its challenges. The future focus of methods may shift towards more integrative, translational tools for massive and heterogeneous collections of data in service of human health. Yet, for such tools to be successful, more research is needed to address ongoing challenges on elucidating the underlying mechanistic basis of diseases. With its computational focus, this collection provides the first steps in this direction.
Please visit the collection at: collections.plos.org/macromolecular
Collection Image Information
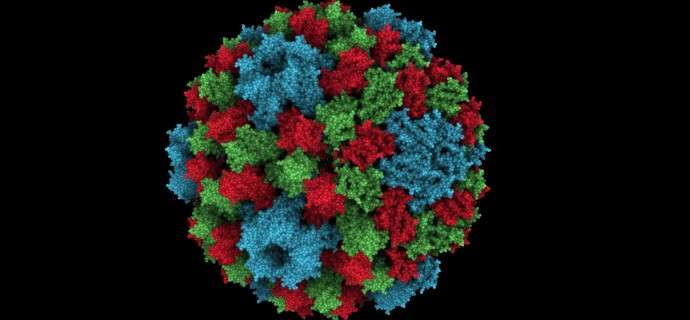
By adding global "ambient occlusion" illumination to a scene, large macromolecular systems look much more natural. Wahle and Wriggers (pcbi.1004516) introduce a novel size-specificity of the shaded features in the Sculptor molecular graphics and modeling software). Shown is a composite of CCMV images (PDB ID 1CWP) with three different scale settings. The illumination can be tuned in real time for pockets and cavities of a user-defined size, making it useful for visualizing large biomolecular systems, comprising e.g. hundreds of thousands of atoms or Mega-Dalton size electron microscopy maps.
More information: Amarda Shehu et al. Computational Methods for Exploration and Analysis of Macromolecular Structure and Dynamics, PLOS Computational Biology (2015). DOI: 10.1371/journal.pcbi.1004585
Journal information: PLoS Computational Biology
Provided by Public Library of Science
This story is republished courtesy of PLOS Blogs: blogs.plos.org.