Researcher developing software to improve dynamic image capture from super-resolution fluorescent microscopes

Techniques such as modifying genes with fluorescent proteins have enabled microscopic images with single-molecule resolution, but MIT researchers are extending techniques designed for acquiring static images to capturing dynamic images with very high resolution in living cells.
"The old mindset of super-resolution imaging is 'I want to make these images that are static pictures of what the cell looks like at some point in time and I want these images to be as sharp as possible,' but for us that's not the most interesting question," explains physics graduate student James Owen Andrews, who works in the laboratory of MIT assistant professor of physics Ibrahim Cissé. "We want to start focusing on dynamics, and so, what all needs to change about the techniques in order to do this ... that's a lot of what I've been thinking about; that and how to analyze the data. It's really exciting working in this field."
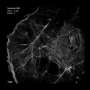
With its emphasis on understanding how RNA polymerase II, an enzyme that reads and copies DNA to create protein-synthesizing RNA molecules, the Cissé Lab uses super-resolution fluorescent microscopes to record polymerase activity in the nucleus of living cells. Recent results point to short-lived clusters of polymerase as the controlling agent in the process. Andrews is developing software to automate the detection and identification of transient clusters in super-resolution data.
Light activates a fluorescent marker in the enzyme, known as a fluorophore, which then emits light of a different color that researchers can track. The super-resolution imaging set up captures images through the microscope onto an EM-CCD (electron-multiplying charge-coupled device), a specialized scientific camera. The EM-CCD has a short exposure time of a few milliseconds, but overall it takes several minutes to get one super-resolution image made from about 10,000 images.
"When you look at these data sets, the difficulty is in actually knowing how long these clusters last. Because of the nature of how the images are acquired, sometimes even a single molecule can look like a cluster, or vice versa," Andrews explains. "Essentially you have a fluorophore—which, for all intents and purposes, is invisible—and then it will be activated and it will be visible for a short time until it burns out. The amount of time that it's on is highly variable, highly random, and so the question is how can you know whether this single molecule which turns on and can last for a couple seconds, how can you distinguish between that single molecule and a cluster of multiple molecules which may last for less than a second or may last for the duration of the video."
At the moment, he says, the most reliable way to do this is by eye. "You can look at images and when the molecules appear in time, and your eye is a lot better at finding patterns than computers are in 2015. The first goal of this software is to simply present data in the most accessible way, which will help you in making the decisions," Andrews says. He is creating modules that read a background image and pop out a window suggesting regions that may have clusters of molecules. The software, written in the MATLAB computing environment, will also attempt to automatically group the data into different independent events.
"We want to fully automate the data analysis for a number of reasons: to make it faster, to make it more reliable, to remove the human element, and to hopefully allow for high-throughput data analysis. We're hoping to be able to start doing studies on hundreds of cells. Where analyzing these cells by hand would take weeks, we hope that a fully automated software package would enable you to start asking questions on hundreds of cells, and have the data analyzed for you by your computer within days, without you having to spend any of your own time on it," he explains. "I have high hopes for it. I think that my most recent version of the software might work but it will take weeks of study before I can say for certain."
If he is successful in completing the software, Andrews says the group expects to publish the software and make it available as a free, editable download. "We want it to be as easy as possible for any lab to be able to study these kinds of events in living cells," Andrews says. "That's really the hope for this lab is to adapt a lot of these techniques to working in living cells so we can study life at its smallest level of organization in real time, in its native context."
Andrews, 25, entered his third year of graduate school this month. He hopes to get his doctorate in the next three to four years. Born in Nashville, Andrews grew up in Marietta, Georgia, and received his bachelor's in physics and chemical and biomolecular engineering from Georgia Institute of Technology in December 2012. Depending on the outcome of his research, his future may lie in academia or in the biotech industry, he says.
Provided by Massachusetts Institute of Technology
This story is republished courtesy of MIT News (web.mit.edu/newsoffice/), a popular site that covers news about MIT research, innovation and teaching.