The life and times of domesticated cheese-making fungi
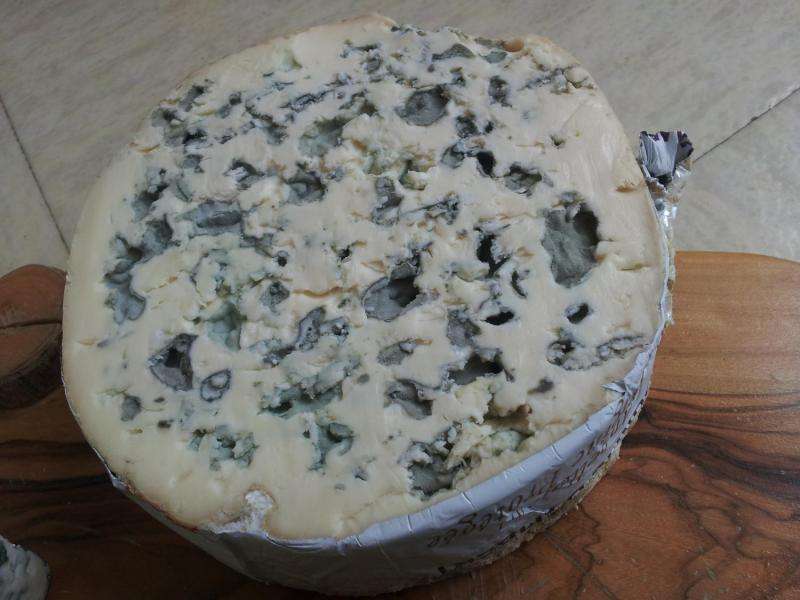

People sure love their cheeses, but scientists have a lot to learn about the fungi responsible for a blue cheese like Roquefort or a soft Camembert. Now researchers reporting in the Cell Press journal Current Biology on September 24 have pieced together the story of those Penicillium fungi, based on the genomes of 14 fungal species. The findings reveal an important role for the horizontal transfer of genes from one species to another in the fungi's ability to trade their freedom in for the domesticated life.
In addition to the evolutionary interest of the findings, they may also lead to improvements in cheese making, the researchers say. The discovery that genes are easily swapped among species also raises concerns for food safety.
"We were able to identify genes that are directly involved in the adaptation to cheese in Penicillium, opening the way for strain improvement, in particular for obtaining fast-growing strains," says Antoine Branca of L'Université Paris-Sud and CNRS. "Our findings, on the other hand, raise concerns about food safety because they suggest that the co-occurrence of different fungal species in the same food product allows genes to be transferred from one species to the other frequently, as it occurred multiple times, for multiple genomic regions."
If food spoilers can readily acquire genes by horizontal gene transfer from cheese-making strains, they may improve their ability to grow in cheeses. That may be a particular concern in the context of fungi that produce toxins.
Branca and co-author Tatiana Giraud realized that cheese-making fungi could make an excellent model for the study of domestication, in part because of the fungi's small genomes. They were also intrigued by the fact that domesticated cheese-making fungi include multiple, distantly related species that have adapted to the same medium, offering striking examples of parallel and convergent evolution. And don't forget that France is famous for its cheeses.
"Cheeses are an emblematic French food, and there is a great diversity of cheese-making fungal species and strains, used for different kinds of cheeses, and thus selected for different traits," Giraud says.
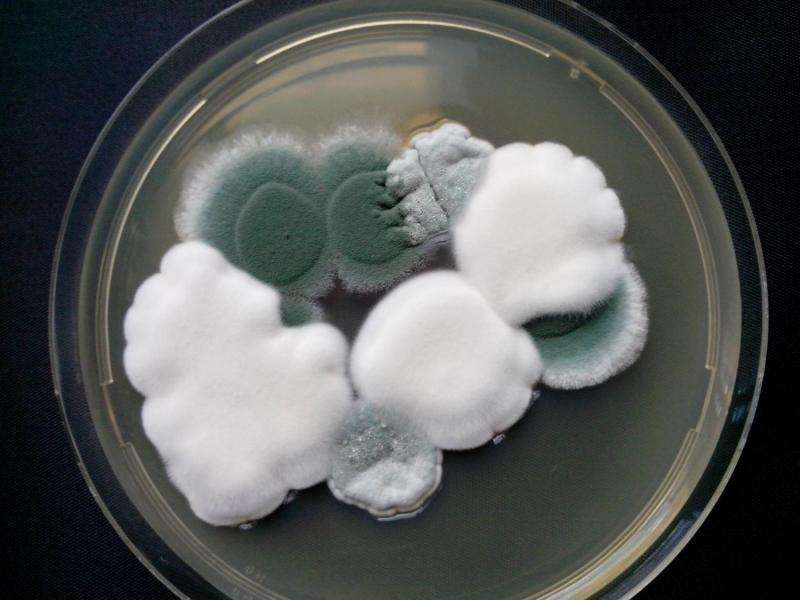
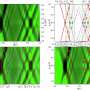
The researchers' comparisons of the Penicillium genomes show that adaptation to cheese was associated with multiple recent horizontal transfers of large genomic regions carrying crucial metabolic genes. They identified seven horizontally transferred regions (HTRs) that were almost identical between distantly related Penicillium species. Two of those regions carried genes involved in the utilization of cheese nutrients or competition among fungal species. Those HTRs endowed fungi with the ability to grow faster and to outcompete other species on cheese.
"It was striking to find such a good match between the putative functions of some genes in the horizontally acquired regions and the function we expected to be essential for metabolizing cheese," Giraud says.
It appears that the fungi's adaptation to life on cheese also comes at a cost. The cheese-making fungi don't fare as well as other species on minimal media. There is plenty left to learn, the researchers say.
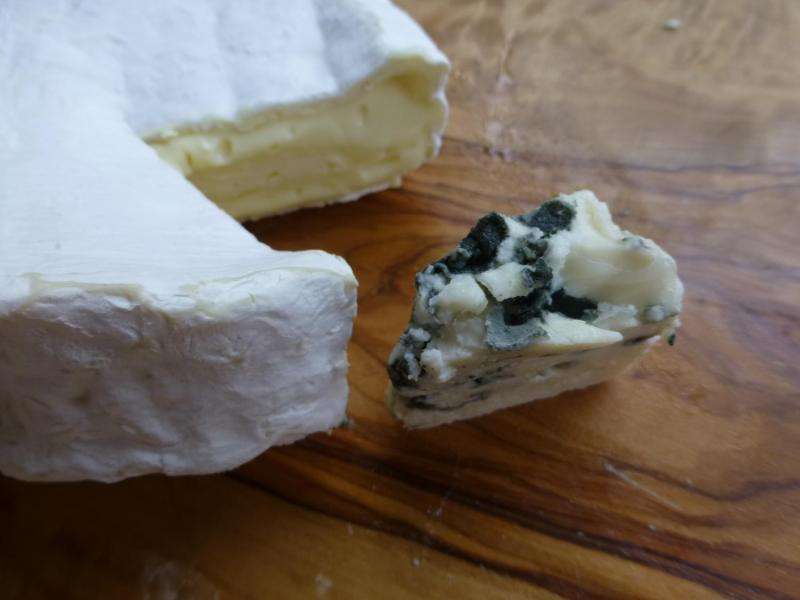

"Strikingly, we still do not know which one is the donor species of the horizontally transferred regions," Branca says.
They don't know where Penicillium roqueforti can be found in nature, either. The fungus has been found on decaying wood, but rarely in natural settings. And yet, the fungus maintains considerable genetic diversity, suggesting that it must exist in relatively large populations.
"If you find a wild blue mold growing outdoors," Branca says, "please let us know!"
More information: Current Biology, Ropars and de la Vega et al.: "Adaptive Horizontal Gene Transfers between Multiple Cheese-Associated Fungi" dx.doi.org/10.1016/j.cub.2015.08.025
Journal information: Current Biology
Provided by Cell Press