Researchers identify unique marker on mom's chromosomes in early embryo
Researchers in the University of Georgia's Regenerative Bioscience Center are visually capturing the first process of chromosome alignment and separation at the beginning of mouse development. The findings could lead to answers to questions concerning the mechanisms leading to birth defects and chromosome instability in cancer cells.
"We've generated a model that is unique in the world," said Rabindranath De La Fuente, an associate professor in the UGA College of Veterinary Medicine. "Because we removed ATRX protein expression only in the oocyte, the female egg cell, we can now study its function at both the cellular and molecular level."
ATRX is a protein that binds to the centromere of all chromosomes in every single cell of the body, but when it malfunctions, chromosomes cannot segregate properly and lose their structural integrity. Using the ATRX protein, the researchers developed a mouse model to learn how an embryo responds to abnormal chromosome segregation.
In the study, published recently in the journal Development, De La Fuente and assistant professor Maria Viveiros, both in the college's department of physiology and pharmacology, have established that stability of a specialized chromosomal domain in an early embryo is absolutely vital for subsequent development and health.
The future goal of this study is to learn about the mechanisms of chromosomal defects, helping to someday reduce the risk of chromosome instability and increase prevention through improving early prenatal care.
There is an urgent need to develop additional non-invasive strategies concerning maternal health, Viveiros said, pointing out the classic example of how folic acid significantly reduced the risk of spina bifida "by the simple recommendation of taking a daily dose of the vitamin folic acid before and during pregnancy.
"With our unique model, by deleting the protein strictly in the female egg, we can begin to understand how maternal proteins help regulate these initial cell divisions during early development."
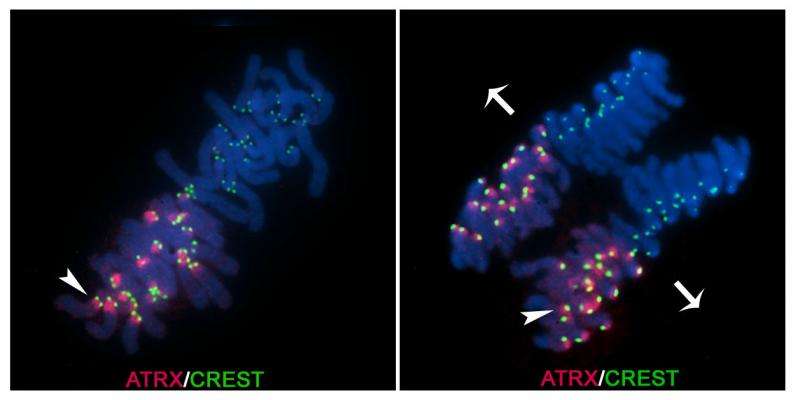
The first image captured by the team shows a mouse oocyte fertilized by sperm, when the maternal and paternal chromosomes come together for the first time to start a new embryo. Through the use of fluorescent markers, the process of how the maternal genome is being regulated can now be studied.
"What's amazing is we can actually visualize that very first division when this cell is going to get half of its chromosomes from mom and the other half from dad," Viveiros said.
They found that ATRX is inherited only from mom's chromosomes.
"That was totally unexpected for us—and the main reason for describing the process as 'epigenetic asymmetry' in the title of our publication," she said.

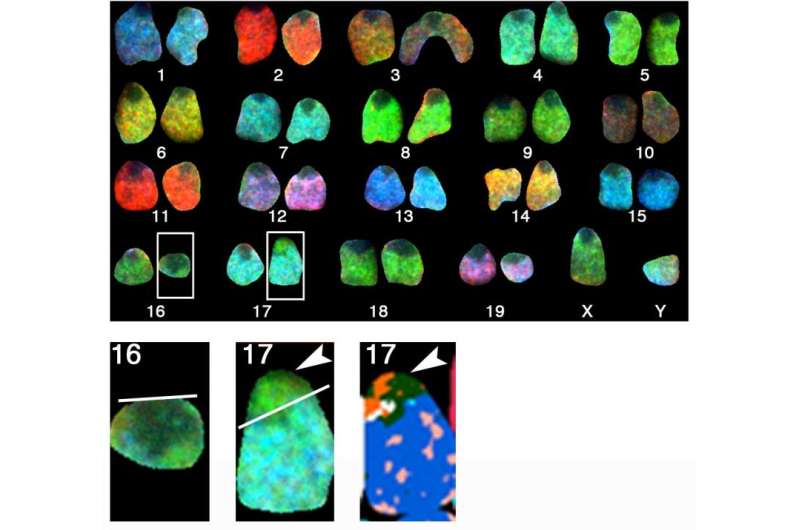
In the second image shown by a computer program that recognizes DNA sequences, in chromosome 16 a piece has gone missing and is now fused with chromosome 17, forming a translocation.
The team has been studying the role of chromatin remodeling proteins in the epigenetic control of chromosome instability for many years, and it's no small task to capture these images and analyze the data. The entire mouse genome is massive and contains billions of base pairs of DNA.
"We've been learning how these proteins work and publishing our results," De La Fuente said, "and at the same time independently in other laboratories around the world oncologists are discovering that ATRX is important to prevent chromosome breaks in tumors. Tumor cells have high rates of genomic instability and are often aneuploid, meaning they inherit the wrong number of chromosomes. This instability is often considered a 'hallmark' for cancer cells. But the mechanisms are not known-we have the model ready to start studying the mechanisms of chromosome instability at the cellular and molecular level."

As to the images presented in the paper, De La Fuente said, "What really keeps us going is a finding like this one, that no one in the world has ever seen before."
More information: "ATRX contributes to epigenetic asymmetry and silencing of major satellite transcripts in the maternal genome of the mouse embryo." Development. 2015 May 15;142(10):1806-17. DOI: 10.1242/dev.118927
Journal information: Development
Provided by University of Georgia