Subsurface discovery sprouts a new branch on the tree of life
Bacteria are enigmatic by nature, minuscule but staggeringly—sometimes alarmingly—ubiquitous. As critical drivers of everything from global biogeochemical cycles to million dollar industries, they are the little cogs that keep our planet going. They even control our brains!
The study of bacteria is hindered because the vast majority of them cannot be cultured or grown in the lab. With the 1977 advent of a molecular technique that enabled culture-independent surveys, microbiologists thought they had a handle on the breadth of microbial diversity on earth.
They were wrong.
Last week a study published in Nature pulled the veil on a branch of the bacterial tree of life that has evaded detection for nearly a century and a half. The study, led by Christopher Brown, who is a PhD candidate in microbial biology at the University of California in Berkeley, used cutting edge genome sequencing and savvy bioinformatics techniques to make this remarkable discovery.
It all started at an abandoned uranium milling site on the banks of the Colorado River in a town called Rifle. This location, contaminated with toxic byproducts of uranium milling, has been a test ground for researchers experimenting on how microbes can be harnessed to bioremediate or clean the environment.
Previously, researchers found that when naturally occurring microbes were supplied with a food source—the simple carbon compound acetate—a fortuitous biochemical reaction ensued. When stimulated to grow, a certain group of bacteria utilized the soluble uranium present in the soil and converted it to an insoluble form. Once insoluble, the uranium would be less apt to flow through groundwater and into the adjacent Colorado River.
Researchers posit microscopic processes like this could be capitalized upon at other contaminated sites to prevent the spread of toxins to drinking water.

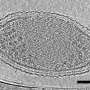
Current research at the Rifle site aims to delve into the ecology of the small but mighty members of this microbial community. In this most recent study, Brown and colleagues went about this by pumping groundwater through a series of filters to capture cells—some of the tiniest free living organisms ever documented.
DNA from these microbial communities was extracted en masse, sequenced into millions of short snippets, then put back together like a jigsaw puzzle—a one so big it necessitated a powerful supercomputer to finish. In the end, this process yielded nearly 800 complete or near-complete microbial genomes.
Even considering the strength of modern high throughput sequencing technology, the recovery of 800 genomes is impressive—especially considering the very first bacterial genome was completed just 20 years ago and required a prodigious entourage of researchers.
Especially surprising about these genomes recovered by Brown and colleagues is that when they tried to place them on the Tree of Life amongst other known bacteria, they didn't fit.
In these ultrasmall organisms, the marker genes typically used in the previously mentioned culture-independent identification experiment were riddled with extra stretches of DNA called insertion sequences.
"We were really surprised to find how diverse these groups are within the bacterial domain, and just how consistently different the organisms within this radiation are from the rest of bacteria," Brown said in a statement.
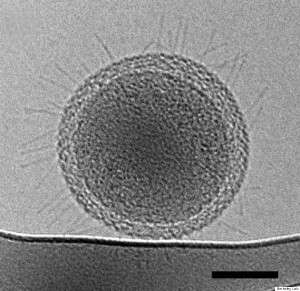
These bacteria cannot be grown in lab cultures. To further complicate things, it is predicted that between 50-100 percent of bacteria in some of the groups they discovered would be completely missed using the standard molecular techniques.
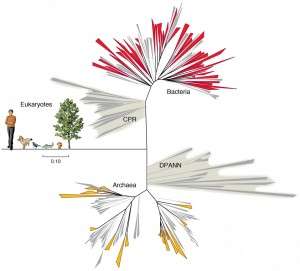
Currently the tree of life is divided into three kingdoms. Bacteria and Archaea are two branches, each composed solely of unicellular organisms. The third kingdom is Eukarya, which encompasses all multicellular life forms and some unicellular microbes as well.
The finding of this paper "represents a substantial modification of the tree of life," corresponding author Jill Banfield said in a statement. "These new major features on the tree of life mean that it probably won't be the simple three-domain view we have now," Banfield said.
These novel bacteria at the Rifle site provide a tantalizing glimpse into microscopic worlds yet to be discovered in other locales. Like the communities at Rifle, other undiscovered bacterial groups could be an untapped resource for beneficial environmental applications.
"People have seen these bacteria in surveys of many different environments all over the planet" Brown said. The next step is to find them.
More information: "Unusual biology across a group comprising more than 15% of domain Bacteria." Nature (2015) DOI: 10.1038/nature14486
Journal information: Nature
Provided by Columbia University