Disabling antibiotic-resistant bacteria
Dreaded bacterial-related diseases have killed untold numbers of humans over the centuries. Today, two million illnesses and nearly 23,000 deaths can be attributed to antibiotic-resistant bacteria throughout the United States, say the Centers for Disease Control and Prevention.
In May, the World Health Assembly took steps to combat antibiotic-resistant bacteria by endorsing a global plan to "improve access to affordable vaccines and address over- and under-nutrition," prompting member states to put plans into action addressing everything from awareness of bacterial resistance to investment in countering antimicrobial resistance.
Antibiotics fight bacteria by targeting enzymes essential to the bacterium's survival, like those involved in rebuilding the bacterium's cellular wall. With many deadly bacteria able to resist antibiotics, scientists must work to design new compounds that target important functions in the bacteria to stop them from multiplying.
As a spinoff from their research aimed at fighting a specific parasite, researchers at the U.S. Department of Energy's Argonne National Laboratory and Brandeis University may have found a way around an infectious bacterium's natural defenses.
Brandeis University researcher Lizbeth Hedstrom initially developed an antimicrobial compound to combat the single-celled parasite Cryptosporidium parvum. The parasite, which infects the small intestine, can cause severe diarrhea in humans who ingest contaminated water. Hedstrom and her team discovered that an enzyme inside this protozoan parasite is similar to a number of enzymes from infectious bacteria.
"Cryptosporidium looks like it got its enzyme gene from a bacterium," Hedstrom said. "Once we knew that, we could make a good guess that bacterial enzymes should be also targeted by the compound we created."
Knowing that both the parasite and a number of infectious bacteria share similar biological processes was the first step.
Hedstrom and her team spent years unlocking Cryptosporidium's structure, cellular processes and key functions. But to test the hypothesis that their antimicrobial compound would work as effectively on bacteria as it does on the parasite, they would have to know just as much about the pathogenic bacteria they wanted to test. So she turned to the University of Chicago's Center for Structural Genomics of Infectious Diseases (CSGID), Argonne's Advanced Protein Characterization Facility (APCF), and the Advanced Photon Source.
"Not only can we produce, define and crystallize proteins, but we can also process thousands of samples very quickly," said Argonne Distinguished Fellow, Computation Institute Senior Fellow and Structural Biology Center Director Andrzej Joachimiak. "With an automated system using robots and state-of-the-art data processing and control software, we can help researchers clone and purify genes quickly."
The CSGID researchers decided to delete one "weak" part of the two-part enzyme, and to their surprise, this made the enzyme much easier to crystallize.
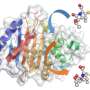
The CSGID and APCF helped Hedstrom quickly define 13 bacterial structures from four different human pathogens. In lab tests, three enzymes were blocked, but the enzyme from the severe diarrhea-causing Vibrio cholerae was unaffected by the compound. Still, images gathered while defining the structure of the bacterial enzyme showed the mechanism by which the enzyme forms and breaks bonds with molecules, and it did so in an unexpected way. This is a rare occurrence that now informs Hedstrom and her team as to the details of process and location of where to target her compound.
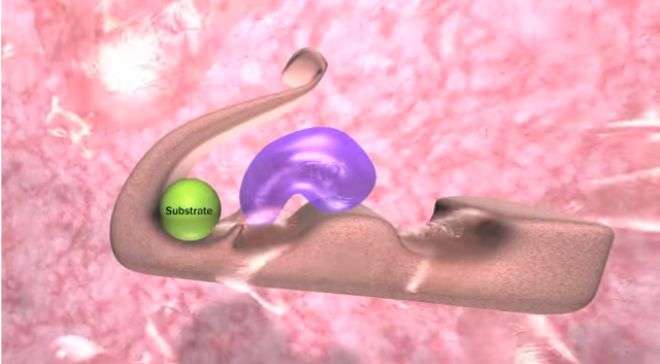
"It is just like a key in a lock," Joachimiak said. "Even though we were not able to inhibit Vibrio cholerae, we weren't able to actually open the door, but now we know how the key we are using should fit the lock."
Hedstrom's next challenge became finding inhibitors that could get into bacteria. Though enzyme targets are very similar, parasites and bacteria differ significantly in how drugs bind to and affect their enzymes. Hedstrom's compounds are designed to diffuse across lipid membranes and into Cryptosporidium, and are therefore very "oily." Unfortunately, bacteria do not take up oily compounds very well, so she needed to make changes to the compound in order to cross the cell wall and enter bacteria. The Argonne structures show exactly where the inhibitors can be modified in order to make sure they fit squarely into the target area without losing the ability to inhibit the function of the enzyme.
Having passed this hurdle, Hedstrom is optimistic about the future of her research and the puzzles she has yet to unlock.
"I am excited to say that we have completed the initial step, which is identifying the compounds," said Hedstrom. "It has been a long process, and we haven't got a drug yet, but we would not be nearly as far along without Andrzej and Argonne's team."
More information: "A Novel Cofactor Binding Mode in Bacterial IMP Dehydrogenases Explains Inhibitor Selectivity." J. Biol. Chem. jbc.M114.619767. First Published on January 8, 2015, DOI: 10.1074/jbc.M114.619767
Provided by Argonne National Laboratory