Researcher develops novel strategy to improve crops and treat diseases

A novel strategy to enhance genome editing promises to increase the efficiency of making genetic improvements in a wide range of organisms, a new study suggests.
The results could help boost applications such as developing better crops and treating genetic diseases in humans, researchers said.
The new strategy is aimed at improving an increasingly popular technique that grew from the recent discovery of a bacterial immune system known as CRISPR-Cas9, according to the study's corresponding author, Yinong Yang, associate professor of plant pathology, Penn State College of Agricultural Sciences.
CRISPR stands for clustered regularly interspaced short palindromic repeats. Yang explained that CRISPR regions of the bacterial genome contain strands of repeating DNA, separated by "spacers" that match the DNA sequences of viruses that have attacked the bacterium or its ancestors.
If attacked again by the same virus, this system allows a bacterium to "remember" and defend against the attacker. The bacterium generates a strand of CRISPR RNA containing a specific spacer sequence that, coupled with a DNA-cutting enzyme known as CRISPR-associated protein nuclease (Cas9), targets the invader and destroys it by slicing its DNA.
"Scientists have discovered that this system can be harnessed as a powerful tool to target and edit almost any DNA sequence in a genome," Yang said. "The CRISPR-Cas technology has broad applications in basic biological research, medicine and agriculture. It is seen as the most important breakthrough in biotechnology so far this century."
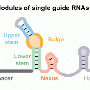
By creating synthetic CRISPR RNA called guide RNA (gRNA) that matches a specific DNA sequence in an organism, scientists can deliver the Cas9 enzyme precisely to the target gene that they want to disable or modify. The process holds promise for precision breeding of crops with desirable traits—such as disease resistance or drought tolerance—and for gene therapy to correct genetic defects that cause human diseases such as sickle-cell anemia and cystic fibrosis.
But Yang noted that many of these applications require the manipulation of more than one gene or target site, creating a need to deploy multiple gRNAs simultaneously. Conventional methods of accomplishing that can be cumbersome or inefficient, typically requiring the stacking of multiple gRNA expression "cassettes" together.
"If you try to express multiple gRNAs by stacking individual expression cassettes, before long you have a DNA fragment that's too large to be cloned efficiently into the vector," he said. "So the need for simultaneous expression of multiple gRNAs is a bottleneck that's preventing us from realizing the full potential of CRISPR-Cas technology."
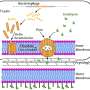
Working with rice as a model system, the researchers overcame this limitation by hijacking the plant's native processing system for transfer RNA, or tRNA. The normal function of tRNA is to deliver amino acids and catalyze bonds between them to synthesize proteins within cells.
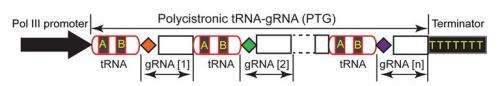
"We designed a synthetic gene with eight tandem repeats of tRNA and gRNA linked together," Yang said. "We found that tRNA-processing enzymes precisely cleaved this architecture at the tRNA-gRNA junction, efficiently releasing multiple gRNAs that can be targeted simultaneously to multiple sites.
"In this study, we targeted eight sites for simultaneous mutation, but in theory, it may be possible to link as many as 20 or 30 tRNAs and gRNAs to enhance multiplex genome editing."
The research team's experiments achieved the desired genetic modifications in rice plants with greater than 90 percent success, which is more efficient than other existing methods, said Yang, who has filed a provisional patent application related to the study. The researchers also used the method successfully on human cell cultures and other plant species such as potato and Arabidopsis.
"Because the tRNA-processing system exists in virtually all organisms, this strategy could be used broadly to boost multiplex genome editing capability of CRISPR-Cas tools," Yang said.
More information: "Boosting CRISPR/Cas9 multiplex editing capability with the endogenous tRNA-processing system." PNAS 2015 ; published ahead of print March 2, 2015, DOI: 10.1073/pnas.1420294112
Journal information: Proceedings of the National Academy of Sciences
Provided by Pennsylvania State University