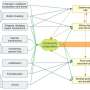
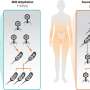
DNA in regions far from target genes governs cell state transitions through a coordinated wave of regulatory activity
By comparing RNA expression patterns across many different tissues and time points, an international research team led by RIKEN scientists has discovered some basic rules of biological regulation. The findings, obtained by mapping the sequence of gene activation when cells undergo a state change, could pave the way for regenerative therapies that modulate cell fate.
Gene expression is activated by two major types of DNA regions: promoters, which are found near the start sites of genes, and enhancers, which are located far from target genes. Scientists had long assumed that both promoters and enhancers act together more or less simultaneously.
Led by RIKEN researchers Piero Carninci and Yoshihide Hayashizaki, a research consortium of more than 100 scientists from 15 countries used a RIKEN-developed technology called CAGE to examine the activity of promoters and enhancers in 19 human cell types and 14 mouse cell types. They analyzed RNA expression in these cells—which included various types of stem cells and fully differentiated lineages—as the cells underwent a variety of changes in response to developmental cues or biological stimuli.
The study showed that, contrary to conventional wisdom, enhancers are the first to be 'turned on', and the switch occurs in the first 15 minutes of a cellular transition. Enhancers then trigger a cascade of regulatory activity, including the production of regulatory genes known as transcription factors that bind and switch on promoters and other regulatory elements. These transcription factors only come into play at the 30–100 minute mark of the cellular time course.
Although the enhancers and transcription factors differed by cell type, these activity patterns were generalizable across the various biological processes studied, which included differentiation, growth and drug treatment. "In cells responding to a wide range of signals, the transcriptional response happens first at enhancers, followed by promoters of transcription factors, followed by promoters of peripheral genes," says lead author Erik Arner from the RIKEN Center for Life Science Technologies.
The study is one of many undertaken by the FANTOM consortium, a research group launched 15 years ago by RIKEN to promote understanding of the function of all expressed RNA transcripts in mouse and human cells. This latest discovery adds to the cellular rulebook and provides scientists with knowledge that could one day assist the development of new therapies. "The more general rules we can discover," says Arner, "the more likely we are to figure out methods to convert any cell type into any other cell type."
More information: Arner, E., Daub, C. O., Vitting-Seerup, K., Andersson, R., Lilje, B., Drablos, F., Lennartsson, A., Rönnerblad, M., Hrydziuszko, O., Vitezic, M. et al. Transcribed enhancers lead waves of coordinated transcription in transitioning mammalian cells. Science 347, 1010−1014 (2015). DOI: 10.1126/science.1259418
Journal information: Science
Provided by RIKEN