Dog waste contaminates our waterways: A new test could reveal how big the problem is
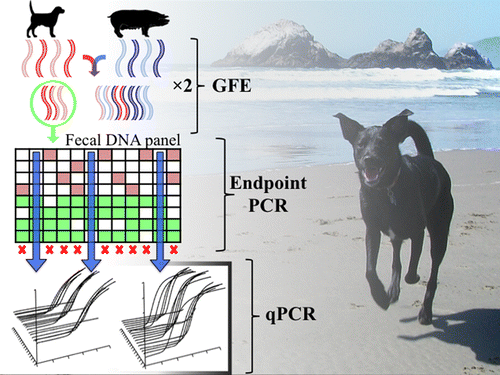
Americans love their dogs, but they don't always love to pick up after them. And that's a problem. Dog feces left on the ground wash into waterways, sometimes carrying bacteria—including antibiotic-resistant strains—that can make people sick. Now scientists have developed a new genetic test to figure out how much dogs are contributing to this health concern, according to a report in the ACS journal Environmental Science & Technology.
Orin C. Shanks, Hyatt C. Green and colleagues explain that our waterways are susceptible to many sources of fecal contamination, including sewage leaks and droppings from farm animals and wildlife. Contamination from dog feces is a concern because it can harbor antibiotic-resistant strains of E. coli and other bacteria and parasites that can infect humans—and there are nearly 70 million domesticated dogs in the U.S.
Scientists have had few tools to determine the extent to which waste from dogs is adding to the pathogens in rivers, lakes and beachfront surf. Current methods look for certain genes from gut bacteria that end up in dog feces. However, this is not foolproof—the microbiota of humans and the canine pets they live with often overlap, making the analysis complicated. So Shanks' team set out to create a more specific test.
The researchers developed a new genetic testing method to specifically detect canine fecal contamination in water. They identified 11 genetic markers that were common among most of the dog samples but missing from the human ones. To determine whether their method would work for real-world monitoring, they sampled storm water from a rain garden where people often walk their dogs. The technique successfully detected some of the same markers they had identified as evidence for canine waste.
More information: "Development of Rapid Canine Fecal Source Identification PCR-Based Assays" Environ. Sci. Technol., Article ASAP. DOI: 10.1021/es502637b
Abstract
The extent to which dogs contribute to aquatic fecal contamination is unknown despite the potential for zoonotic transfer of harmful human pathogens. We used genome fragment enrichment (GFE) to identify novel nonribosomal microbial genetic markers potentially useful for detecting dog fecal contamination with PCR-based methods in environmental samples. Of the 679 sequences obtained from GFE, we used 84 for the development of PCR assays targeting putative canine-associated genetic markers. Twelve genetic markers were shown to be prevalent among dog fecal samples and were rarely found in other animals. Three assays, DG3, DG37, and DG72, performed best in terms of specificity and sensitivity and were used for the development of SYBR Green and TaqMan quantitative PCR (qPCR) assays. qPCR analysis of 244 fecal samples collected from a wide geographic range indicated that marker concentrations were below limits of detection in noncanine hosts. As a proof-of-concept, these markers were detected in urban stormwater samples, suggesting a future application of newly developed methods for water quality monitoring.
Journal information: Environmental Science & Technology
Provided by American Chemical Society