How to tell good stem cells from the bad
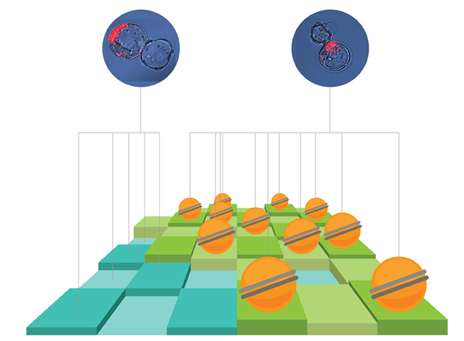
The promise of embryonic stem cell research has been thwarted by an inability to answer a simple question: How do you know a good stem cell from a bad one?
Yale researchers report in the Sept. 4 issue of the journal Cell Stem Cell that they have found a marker that predicts which batch of personalized stem cells will develop into a variety of tissue types and which will develop into unusable placental or tumor-like tissues.
Scientists have been unable to capitalize on revolutionary findings in 2006 that adult cells could be made young again with the simple introduction of four factors. Hopes were raised that doctors would soon have access to unlimited supplies of a patient's own iPSCs—induced pluripotent stem cells—that could be used to repair many types of tissue damage. However, efforts to direct these cells to therapeutic goals have proved difficult. Many attempts to use cells clinically have failed because they form tumors instead of the desired tissue.
The team of Yale Stem Cell Center researchers led by senior author Andrew Xiao identified a variant histone—a protein that helps package DNA—which can predict the developmental path of iPSC cells in mice. An accompanying paper in the same journal by researchers at the Whitehead Institute at MIT and Hebrew University in Israel also identifies at different marker that also appears to predict stem cell fate.
"The trend is to raise the standards and quality very high, so we can think about using these cells in clinic," Xiao said. "With our assay, we have a reliable molecular marker that can tell what is a good cell and what is a bad one."
Journal information: Cell Stem Cell
Provided by Yale University