A key component of cell division comes to light
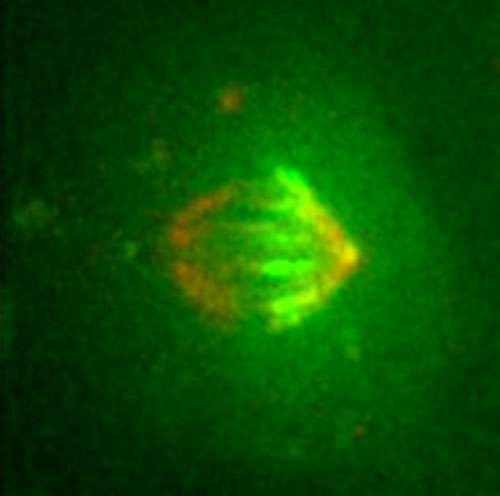
The division of a cell in two requires the assembly of the mitotic spindle, an extremely complex structure, which is the result of the coordinated action of a multitude of proteins and a finely tuned balance of their activities. A large part of the time that a cell requires to divide is devoted to assembling the mitotic spindle, which, superficially, resembles a ball of thread with the shape of a rugby ball. The most abundant components of the spindle are the microtubules. "By labelling the ends of thousands of these fine filaments, which are indispensable and extremely dynamic and variable, we have finally been able to follow their distribution and movement during the assembly of the mitotic spindle," explains Jens Lüders, a cell biologist from the Institute for Research in Biomedicine (IRB Barcelona). The breakthrough appeared yesterday in the advanced online edition of the journal Nature Cell Biology.
"For more than 10 years we have been able to track only the growing ends of microtubules but not the starting points. As a result, we lacked essential information in order to understand the dynamic architecture of the mitotic spindle and how it contributes to cell division," says Lüders. Headed by the German scientist who runs the Microtubule Organisation Group at IRB Barcelona, the study carries only two names, his own and that of the French researcher Nicolas Lecland, first author, who completed his PhD at IRB Barcelona through a "la Caixa" fellowship.
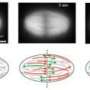
The scientists have demonstrated that the protein γ-tubulin localizes at the starting points of the microtubule filaments and is relatively stably associated with these structures. Using a version of γ-tubulin that carries a fluorescent label activated by laser light, the researchers were able to follow the movement of the starting points of microtubules within mitotic spindles by filming dividing human cells. The Advanced Digital Microscopy Facility, a joint IRB Barcelona-Barcelona Science Park Facility run by the IRB physicist Julien Colombelli, has been crucial for setting up the technology required. "The success of this study is also the result of the technical know-how and cutting-edge technology available, without which we would never have been able to tackle this project," emphasizes Lüders.
The researchers describe for the first time where most microtubules form inside the mitotic spindle, how they develop, and how their starting points are transported—with the help of three motor proteins—to opposite poles of the spindle, where they attach. Simultaneous to this process, the opposite ends of the filaments extend towards the cell centre, where they interact with chromosomes. When the spindle is finally assembled, the microtubules pull the chromosomes to opposite poles and initiate the physical division of the cell. "We now have a more complete understanding of how the spindle assembles and functions and can use our novel marker for testing old and new hypotheses about underlying mechanisms," says the scientist.
A new tool to study cancer
In addition, the breakthrough paves the way to better understanding the mode of action of drugs that inhibit microtubules and that are used in chemotherapy. These kinds of drugs impede the mitotic spindle, thus preventing cell division and interfering with tumour growth.
In spite of the many years of clinical success of these treatments against cancer, little is known about how they impair spindle architecture and function. Although these drugs are highly efficient, they do not show the specificity desirable as they also affect healthy dividing cells. In addition, they affect non-dividing cells such as neurons, in which microtubules also have important functions. "A better understanding of the differences in spindle organisation between cancer and healthy cells and how they respond to microtubule-targeted drugs is essential in order to optimise treatments, for example by identifying more specific drugs or new targets. This tool could be useful to achieve these objectives," states the researcher.
More information: The dynamics of microtubule minus ends in the human mitotic spindle, Nicolas Lecland and Jens Lüders, Nature Cell Biology (2014) dx.doi.org/10.1038/ncb2996
Journal information: Nature Cell Biology