Defining function from "genomic dark matter"
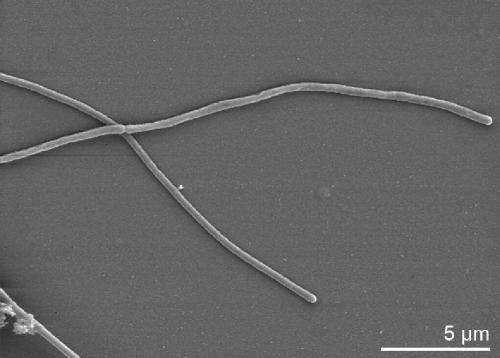

Functional annotation allowed researchers to identify biomass-degrading enzymes in the 35 percent of genes in a genome that are considered "genomic dark matter."
Identifying 17 putative biomass-degrading cellulases from the content of more than 5,500 microbial genomes is of use to bioenergy researchers working on efficiently converting plant biomass into advanced biofuels.
Bioenergy proponents have long held that the amount of plant biomass available in the United States could supplement and even substantially replace the use of food crops for biofuels. The challenge however, has been converting the plant biomass efficiently and cost-effectively to make the switch from gasoline to alternative fuels more feasible. To help meet this challenge, researchers have been working on identifying microbial enzymes that can break down plant biomass. One such study led by the U.S. Department of Energy Joint Genome Institute (DOE JGI) harnessed massive-scale DNA sequencing to identify nearly 30,000 biomass-degrading genes from microbes found in the cow rumen. Testing a small fraction of these genes revealed that more than half of them were in fact capable of breaking down plant biomass, suggesting that a significant number of these putative genes would have the same capability.
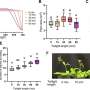
Now, a recent study published online in the April 12, 2014 issue of Biotechnology and Bioengineering focuses on the computational challenges of identifying novel biomass-degrading genes from existing microbial genomes. As much as a third of the genes identified in most genomes currently remain unknown, hindered in part by the inability of existing algorithms to assign functions to these genes and proteins. A solution proposed by the researchers considers the genomic context and sequence similarity of these unknown genes to already-known cellulases. To test this approach, they screened more than 5,500 microbial genomes on the Integrated Microbial Genomes (IMG) database maintained by the DOE JGI and identified 56 "hypothetical proteins" without assigned function. All cellulolytic proteins were detected on genomes that had been sequenced as part of the Genomic Encyclopedia of Bacteria and Archaea project. Seventeen of these candidate cellulases were then directly tested for their cellulolytic activity with 11 (65%) showing activity. The team observed an additional 45 cellulase candidates that were identified but not tested. Using the bioinformatics tools available on the IMG site, they added, would allow researchers to build on the work they started.
"In summary," the researchers wrote, "this work provides initial insights into the cellulolytic capability of 'genomic dark matter' that contributes up to ~1/3 of all genes in a genome…. The set of new biomass-degrading genes and corresponding proteins identified in this study will provide a valuable extension of the overall catalogue of carbohydrate-active genes and proteins currently available." This approach may also have utility in exploring genomic dark matter for additional enzymes of relevance to DOE interests in bioenergy and the environment.
More information: Piao H et al." Identification of novel biomass-degrading enzymes from genomic dark matter: Populating genomic sequence space with functional annotation." Biotechnol Bioeng. 2014 Apr 12. DOI: 10.1002/bit.25250. [Epub ahead of print]
Journal information: Biotechnology and Bioengineering
Provided by US Department of Energy