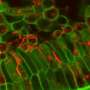
Researchers annotate genome of the smallest known fungal plant pathogen

Researchers sequenced and analyzed the genome of Mixia osmundea, the smallest fungal plant pathogen (13.6 million bases) to date, to provide insight into its mode of pathogenicity and reproductive biology.
Aside from learning how the fungal pathogen reproduces, genome annotation revealed its capabilities in breaking down plant cell wall components, which is of interest to bioenergy researchers.
With a kingdom of more than a million species, fungi thrive in diverse ecological niches, play roles in plant health, and are considered to have a whole host of untapped, as-yet unknown applications. At the Department of Energy Joint Genome Institute (DOE JGI), fungal researchers are involved in several worldwide collaborations to learn more about fungi, and how they can be harnessed for roles such as improving the health of candidate biomass feedstocks for biofuels development.
As part of the work toward illuminating the fungal tree of life, a study published in the April 2014 issue of New Phytologist focused on a tiny plant pathogen called Mixia osmundea, a member of the Pucciniomycotina family that has had few species sequenced to date. Though it was first described 100 years ago, the fungus is rarely seen and has only been isolated on the fronds of two fern species in Japan, Taiwan and America. The genome was sequenced and annotated and the data have been made available on the DOE JGI's fungal portal MycoCosm.
During the team's analysis of the fungus, they identified several carbohydrate-active enzymes that indicate M. osmundea's capabilities in breaking down plant mass, as expected of the plant pathogen. They also found multiple copies of an enzyme that suggests the fungus is "especially efficient" at breaking down a particular compound in plant cell walls. However, they did not find genes that would indicate the fungus can break down xylan, or convert cellobiose into glucose. "Therefore," the team wrote, "although M. osmundae seems to possess a set of enzymes that can be used to break down cellulose, it lacks the enzyme sets necessary for depolymerizing it to simple sugars."
More information: Toome M et al. "Genome sequencing provides insight into the reproductive biology, nutritional mode and ploidy of the fern pathogen Mixia osmundae." New Phytol. 2014 Apr;202(2):554-64. DOI: 10.1111/nph.12653
Journal information: New Phytologist
Provided by US Department of Energy