Antibiotic 'smart bomb' can target specific strains of bacteria
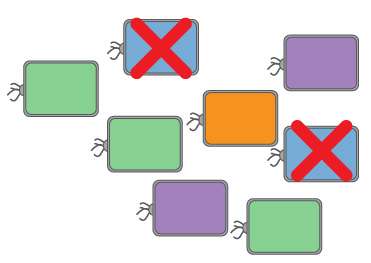

Researchers from North Carolina State University have developed a de facto antibiotic "smart bomb" that can identify specific strains of bacteria and sever their DNA, eliminating the infection. The technique offers a potential approach to treat infections by multi-drug resistant bacteria.
"Conventional antibiotic treatments kill both 'good' and 'bad' bacteria, leading to unintended consequences, such as opportunistic infections," says Dr. Chase Beisel, an assistant professor of chemical and biomolecular engineering at NC State and senior author of a paper describing the work. "What we've shown in this new work is that it is possible to selectively remove specific strains of bacteria without affecting populations of good bacteria."
The new approach works by taking advantage of a part of an immune system present in many bacteria called the CRISPR-Cas system. The CRISPR-Cas system protects bacteria from invaders such as viruses by creating small strands of RNA called CRISPR RNAs, which match DNA sequences specific to a given invader. When those CRISPR RNAs find a match, they unleash Cas proteins that cut the DNA.
The NC State researchers have demonstrated that designing CRISPR RNAs to target DNA sequences in the bacteria themselves causes bacterial suicide, as a bacterium's CRISPR-Cas system attacks its own DNA.
"In lab testing, we found that this approach removes the targeted bacteria," Beisel says. "We're still trying to understand precisely how severing the DNA leads to elimination of the bacteria. However, we're encouraged by the ease in specifically targeting different bacteria and the potency of elimination."
The researchers tested the approach in controlled cultures with different combinations of bacteria present, and were able to eliminate only the targeted strain. "For example, we were able to eliminate Salmonella in a culture without affecting good bacteria normally found in the digestive tract," Beisel says.
The researchers were also able to demonstrate the precision of the technique by eliminating one strain of a species, but not another strain of the same species which shares 99 percent of the same DNA.
Another benefit of the approach, Beisel says, is that "by targeting specific DNA strands through the CRISPR-Cas system, we're able to bypass the mechanisms underlying the many examples of antibiotic resistance."
The researchers are currently working to develop effective methods for delivering the CRISPR RNAs in clinical settings.
"This sets the stage for next-generation antibiotics using programmable CRISPR-Cas systems," says Dr. Rodolphe Barrangou, an associate professor of food, bioprocessing and nutrition sciences at NC State and co-author of the manuscript.
More information: The paper, "Programmable removal of bacterial strains by use of genome-targeting CRISPR-Cas systems," is published online in the journal mBio. mbio.asm.org/content/5/1/e00928-13
Abstract
CRISPR (clustered regularly interspaced short palindromic repeats)-Cas (CRISPR-associated) systems in bacteria and archaea employ CRISPR RNAs to specifically recognize the complementary DNA of foreign invaders, leading to sequence-specific cleavage or degradation of the target DNA. Recent work has shown that the accidental or intentional targeting of the bacterial genome is cytotoxic and can lead to cell death. Here, we demonstrate that genome targeting with CRISPR-Cas systems can be employed for the sequence-specific and titratable removal of individual bacterial strains and species. Using the type I-E CRISPR-Cas system in Escherichia coli as a model, we found that this effect could be elicited using native or imported systems and was similarly potent regardless of the genomic location, strand, or transcriptional activity of the target sequence. Furthermore, the specificity of targeting with CRISPR RNAs could readily distinguish between even highly similar strains in pure or mixed cultures. Finally, varying the collection of delivered CRISPR RNAs could quantitatively control the relative number of individual strains within a mixed culture. Critically, the observed selectivity and programmability of bacterial removal would be virtually impossible with traditional antibiotics, bacteriophages, selectable markers, or tailored growth conditions. Once delivery challenges are addressed, we envision that this approach could offer a novel means to quantitatively control the composition of environmental and industrial microbial consortia and may open new avenues for the development of "smart" antibiotics that circumvent multidrug resistance and differentiate between pathogenic and beneficial microorganisms.
Journal information: mBio
Provided by North Carolina State University