Hijacked protein may lead to new therapeutic interventions
(Phys.org) —Researchers hope to hijack a natural process called RNA interference to block the production of proteins linked to disease and treat medical conditions for which conventional drugs do not work, including cancer, heart disease, HIV and Parkinson's disease.
Scientists working at SLAC's Stanford Synchrotron Radiation Lightsource (SSRL) recently took a significant step in that direction: They used X-rays to shed light on a key component of the RNA interference machinery in human cells – a protein called Argonaute2. The study, published in the journal Science, provides the first complete structural picture of this important protein.
Armed with this information, the researchers hope to design molecules that would guide Argonaute2 to specific targets in the cell and fight disease.
"This approach is highly adaptable and has the potential of targeting a wide variety of diseases," explains Ian MacRae of The Scripps Research Institute in La Jolla, who performed the experiments with Scripps colleague Nicole Schirle. "Some of these interventions are currently being tested in clinical trials."
Hijacking a natural process
Conventional drugs consist of small molecules that bind to disease-linked proteins, block their functions and suppress the disease. Unfortunately, most of these proteins do not have proper target sites for drugs and are considered "non-druggable."
RNA interference offers an alternative: Rather than attacking a protein directly, it destroys messenger RNA molecules that carry instructions for making the protein. To do this, researchers need to learn how cells normally carry out this function to get rid of unwanted messenger RNA.
One of the key players is Argonaute2. "It silences messenger RNA in different tissues as part of normal development," says MacRae. It does this by first binding to another RNA molecule known as guide RNA, which leads it to the messenger RNA that is targeted for degradation.
In principle, researchers can program Argonaute2 to destroy any desired messenger RNA by attaching it to the appropriate guide RNA. However, designing those guides requires detailed knowledge of Argonaute2's structure and how it interacts with both guide and messenger RNAs.
Implications for designing therapeutics
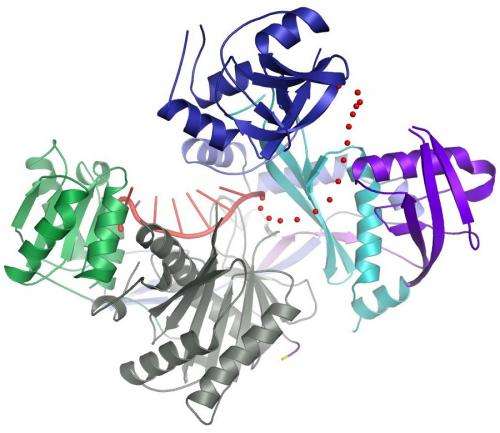
Schirle and MacRae have now determined the structure of the human Argonaute2 protein for the first time. Previously, only parts of the human protein and similar full-length proteins from bacteria had been studied.
The structure revealed in detail how one part of the guide RNA binds to Argonaute2, leaving another part free to bind messenger RNA. Moreover, a segment of the guide RNA forms a helix, which helps it locate and bind to messenger RNA. The scientists also showed how Argonaute2 interacts with other proteins that accelerate the decay of messenger RNA.
The results will likely impact developments in the field of RNA interference therapeutics. To target particular messenger RNAs, companies now produce large numbers of guide RNAs with different random modifications and screen them to determine which ones work best. Now, says MacRae, a more directed and efficient approach to the development of RNA interference therapeutics may be within reach.
More information: www.sciencemag.org/content/336/6084/1037
Journal information: Science
Provided by SLAC National Accelerator Laboratory