Geneticists show COPIA-R7 transposon enhances immunity of its host against pathogenic microorganism
(Phys.org) —Transposons are DNA elements that can multiply and change their location within an organism's genome. Discovered in the 1940s, for years they were thought to be unimportant and were called "junk DNA." Also referred to as transposable elements and jumping genes, they are snippets of "selfish DNA" that spread in their host genomes serving no other biological purpose but their own existence.
Now Tokuji Tsuchiya and Thomas Eulgem, geneticists at the University of California, Riverside, challenge that understanding. They report online this week in the Proceedings of the National Academy of Sciences that they have discovered a transposon that benefits its host organisms.
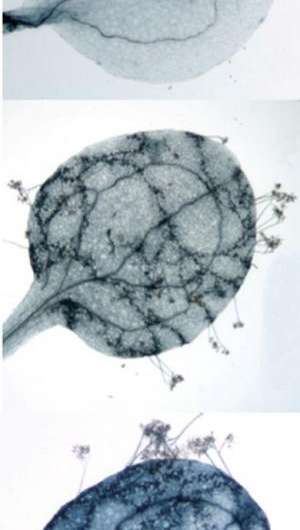
Working on the model plant Arabidopsis, they found that the COPIA-R7 transposon, which has jumped into the plant disease resistance gene RPP7, enhances the immunity of its host against a pathogenic microorganism that is representative of a large group of fungus-like parasites that cause various detrimental plant diseases.
"We provide a new example for an 'adaptive transposon insertion' event—transposon insertions that can have beneficial effects for their respective host organisms—and uncover the mechanistic basis of its beneficial effects for plants," said Thomas Eulgem, an associate professor of plant cell biology and the senior author of the research paper. "While it has been known for a while that transposon insertions can have positive effects for their respective host organisms and accelerate evolution of their hosts, cases of such adaptive transposon insertions have been rarely documented and are, so far, poorly understood."
The COPIA-R7 transposon affects RPP7 by interfering with the latter's epigenetic code. In contrast to the well known 4-letter genetic DNA code, which provides instructions for the synthesis of proteins, the "epigenetic code" defines the activity states of genes and determines to what extent their genetic information is utilized. Eulgem explained that the transposition of transposons is typically inhibited by epigenetic silencing signals associated with their DNA. Such epigenetic signals are like molecular "flags" or "tags" that are attached to special proteins, around which DNA is wrapped.
A type of molecular flag, referred to as H3K9me2, prohibits transposons from being active and jumping in their host genomes.
"An exciting aspect of our work is that H3K9me2 signals associated with COPIA-R7 have acquired a completely new meaning in RPP7 and promote the activity of this disease-resistance gene," said Eulgem, a member of UC Riverside's Center for Plant Cell Biology. "By modulating levels of this silencing signal in RPP7, plants can adjust the activity of this disease resistance gene.
"Silencing of transposon activity is a complex process that is based on the interplay between different types of epigenetic signals," Eulgem continued. "Typically H3K9me2 is of critical importance for transposon silencing. However, we found H3K9me2 is not important for COPIA-R7 silencing, perhaps because this type of epigenetic signal has acquired a different function within the RPP7 gene. While we found H3K9me2 to promote RPP7 activity, it seems to have lost its function for COPIA-R7 silencing."
Arabidopsis plants use H3K9me2-mediated messenger RNA processing to accurately set RPP7 activity to precisely defined levels. In principle, scientists interested in crop improvement can now use the UCR discovery to design new types of molecular switches based on H3K9me2-mediated messenger RNA processing. Using standard molecular biological methods, transposon sequences that are naturally associated with this epigenetic signal can be inserted into suitable genes and thereby alter the activity levels of these genes.
"Our results are critical for the basic understanding of how transposons can affect the evolution of their hosts—something not well understood at this time," said Tokuji Tsuchiya, the first author of the research paper and an assistant specialist in Eulgem's lab. "Besides this impact on basic research, the epigenetic mechanism we discovered can possibly be utilized for biotechnological crop improvement. In principle, the switch mechanism we discovered can be applied to all crop species that can be genetically modified."
Next, Eulgem plans to expand his lab's research to how plants use the modulation of H3K9me2 levels at COPIA-R7 to dynamically adjust RPP7 activity when they are attacked by a pathogenic microorganism and to explore if this mechanism also applies to additional genes.
"It would make sense to assume that at other transposons, H3K9me2 levels are also modulated during immune responses and that this epigenetic mark affects the activity of other genes that are important for plant immunity," Eulgem said. "If this is true, we have uncovered a completely new genetic—or epigenetic—mechanism that allows plants to sense that they are under pathogen attack and to initiate appropriate immune responses."
More information: www.pnas.org/content/early/201 … /1312545110.abstract
Journal information: Proceedings of the National Academy of Sciences
Provided by University of California - Riverside