April 29, 2013 report
Bioengineering team creates self-forming tetrahedron protein

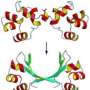
(Phys.org) —A combined team of researchers from the U.S. and Slovenia has succeeded in creating "origami" type proteins that assemble themselves into three dimensional shapes. As a proof of concept, the team created, as they describe in their paper published in the journal Nature Chemical Biology, a protein with coils that self-formed a true three dimensional tetrahedron.
For several years DNA "origami" has been in the news as bio-nanotechnologists have created strands that self-form into boxes, smiley faces etc. Since that time, the attention of many such researchers has switched over to doing the same, or better, with proteins—the thinking has been that such proteins would prove useful for medical or nano-engineering projects. Up until now, however, doing so has been slow going.
Proteins are long chains of amino acids (polypeptides) that, in nature, exist in many different shapes—their purpose is to perform services inside of cells to help keep them functioning properly. One of the most common natural protein structures is a coil—it like other protein structures holds its shape due to attraction between amino acids—the order of the acids in the chain helps determine which overall shape results. The researchers used this knowledge to create proteins that self-formed into new shapes.
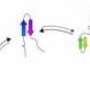
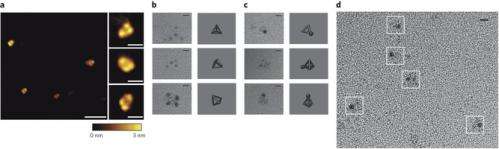
To make the tetrahedron, the researchers began with proteins that already self-form into coils—in this case, 12 of them to represent each of the edges of the desired shape. Then, they added in a different type of amino acid to serve as both linker and joint holding the 12 coils together as a single chain. To bring the protein into existence, the team used genetically altered Escherichia coli as the host—it synthesized the protein, which then self-formed into the tetrahedron as planned—the first such structure to be created in a lab.
Creating customized three dimensional protein structures may soon lead to new ways to deliver medicine—molecules could be embedded inside of them for, example, and released by another chemical once it arrives at the right destination. Nanotechnologists are also interested in creating specific protein structures because of their possible use in creating bio-data-storage devices or other useful electronic materials.
More information: Design of a single-chain polypeptide tetrahedron assembled from coiled-coil segments, Nature Chemical Biology (2013) doi:10.1038/nchembio.1248
Abstract
Protein structures evolved through a complex interplay of cooperative interactions, and it is still very challenging to design new protein folds de novo. Here we present a strategy to design self-assembling polypeptide nanostructured polyhedra based on modularization using orthogonal dimerizing segments. We designed and experimentally demonstrated the formation of the tetrahedron that self-assembles from a single polypeptide chain comprising 12 concatenated coiled coil–forming segments separated by flexible peptide hinges. The path of the polypeptide chain is guided by a defined order of segments that traverse each of the six edges of the tetrahedron exactly twice, forming coiled-coil dimers with their corresponding partners. The coincidence of the polypeptide termini in the same vertex is demonstrated by reconstituting a split fluorescent protein in the polypeptide with the correct tetrahedral topology. Polypeptides with a deleted or scrambled segment order fail to self-assemble correctly. This design platform provides a foundation for constructing new topological polypeptide folds based on the set of orthogonal interacting polypeptide segments.
Journal information: Nature Chemical Biology
© 2013 Phys.org