Shedding light on chemistry with a biological twist

(Phys.org) —Many of life's processes rely on light to trigger a chemical change. Photosynthesis, vision, the movement of light-seeking or light-avoiding bacteria, for instance, all exploit photochemistry. Discovering exactly how living things absorb and convert light energy into a form that can change the molecules involved in such processes would not only help scientists understand them but could lead to ways to mimic such processes for more efficient solar energy conversion, for instance. A clearer understanding of how light can drive biological processes has emerged from x-ray diffraction studies carried out on beamlines at the U.S. Department of Energy Office of Science's Advanced Photon Source (APS) at Argonne, and the European Synchrotron Radiation Facility (ESRF). This work will help science shed a brighter light on some of life's most critical processes.
At the heart of these processes are photoactive protein molecules. Under the right conditions, when light shines on them, specific chemical bonds within the light-absorbing part of the protein can undergo a chemical change called isomerization.
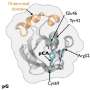
For example, photoactive yellow protein (PYP) is a light receptor in the bacterium Halorhodospira halophila. When PYP is activated by light it triggers a cascade of biochemical reactions that allow the bacterium to move away from the light, a process known as negative phototaxis. At the heart of PYP is a color center, or chromophore, that absorbs the required photons of light and in so doing undergoes a chemical transformation—a trans to cis isomerization. This chemical flip in turn changes the behavior of PYP itself and signals the start of that cascade.
The use of the word "flip" belies a much more complicated process than a straightforward transformation of the chromophore, as the researchers from the Institute for Basic Science (Republic of Korea), KAIST (Republic of Korea), The University of Wisconsin–Milwaukee, and The University of Chicago discovered. They point out that the complexity of any protein discounts the simplistic one-bond flip of the kind that might take place in simple laboratory chemicals. Instead, they suggest that sophisticated chemical choreography must take place to allow the flip to occur without disrupting the overall shape and volume of the protein.
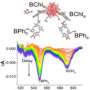
The team utilized time-resolved Laue diffraction x-ray crystallography at the BioCARS 14-ID-B beamline at the APS and the ESRF ID09B beamline to build up a frame-by-frame picture of the changes occurring every 100 ps, at 1.6-Å resolution, as the chromophore absorbs light and the photochemical reactions in PYP begin.
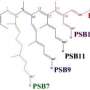
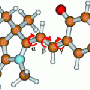
The PYP chromophore is p-coumaric acid, a molecule widespread in nature and present in several edible plants including tomatoes and garlic, as well as the bacterium on which the research team has focused. This molecule contains a double bond between two central carbon atoms in its structure. Such bonds are usually amenable to the kind of transformation that involves a flip from a trans to a cis state where chemical groups linked by the double bond shift from both being on opposite sides to both being on the same side. In the trans state, p-coumaric acid gives rise to the yellow color of PYP, but when it changes to cis, having absorbed a photon, its color is lost and the light energy is transformed into the required physical change in the protein.
Whereas in the laboratory the trans-to-cis flip can occur in a straightforward manner, the team's crystallographic studies of the transformation as it occurs within the protein reveal that an intermediate is involved. This intermediate is a highly strained molecule in which the chemical groups on either side of the double bond in p-coumaric acid are forced to align from their original positions on either side before the molecule releases its pent up energy to generate the cis form with both groups on the same side.
Even this apparently simple release is complicated by the existence of two reaction paths: the "hula-twist" and the "bicycle-pedal." These distinct reaction routes allow the groups adjacent to the double bond either to squirm into the cis position, in the former case, or to rotate around the bond like rotating bicycle pedals. The team has used a mutant version of the protein to control which of these two pathways is taken. A mutation in PYP known as E46Q weakens the connection between p-coumaric acid and the protein itself and this seems to preclude the process from taking the bicycle pedal route.
These findings will help scientists better interpret spectroscopic data from this and similar systems as well as improving their computational models all shedding more light on some of life's most critical processes.
More information: Jung, Y. et al. Volume-conserving trans–cis isomerization pathways in photoactive yellow protein visualized by picosecond X-ray crystallography, Nat. Chem. 5, 212 (March 2013). DOI:10.1038/NCHEM.1565
Provided by Argonne National Laboratory