Study reveals complex process behind sea sponge's toxicity

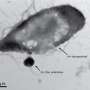
(Phys.org)—The solution to a biochemical puzzle over the molecular make-up of a coral reef sea sponge (Theonella swinhoei) has revealed the origin of its extremely toxic agents.
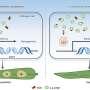
A team of international scientists, including academics at The University of Nottingham, have discovered that polytheonamides— highly toxic and chemically complex compounds which are key to the sponge's defences—are produced by the organism in disguise.
And in a final twist they found that the polytheonamides are not even made by the sponge itself but by a bacterium living with it in symbiosis.
Their discoveries, published on the Science Express website last week, could have implications for human medicine too—the highly effective nature of the sponge's toxins, and their means of production, could be the starting point for new antibiotics and anti-cancer therapies.
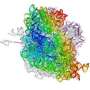
Polytheonamides belong to a class of molecules known as peptides, which are polymers built from amino acid units. Peptides and proteins are normally made by the ribosome, a molecular machine which translates the genetic code, originating from DNA, into the amino acid code required to assemble proteins.
The ribosome uses only 22 amino acids in its 'vocabulary' and because polytheonamides contain many other unusual amino acids they have previously been classified as non-ribosomal peptides, which are made by enzymes like the vast majority of natural products and not read directly from the genetic code.
The international team, which also included scientists from Bonn and Tokyo, has demonstrated that, despite their highly unusual amino acid content, the polytheonamides are in fact ribosomal products in disguise which have undergone an unprecedented degree of modification in their early formation.
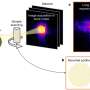
The experts believe that toxicity lies at the heart of this baffling but ingenious piece of bioengineering. The modifications change the three dimensional shape of the peptide into a helix that can insert into a cell membrane to make an ion channel which disrupts the membrane's electrical potential and kills the cell.
Dr Neil Oldham, of The University of Nottingham's School of Chemistry, said: "Understanding how these molecules are produced has been a real challenge: they are almost unrecognisable as products of the ribosome."
"Cell kill activity is the starting point for many anti-cancer and antibiotic strategies. Having direct control over the structure of these molecules may allow modulation of activity through 'designed biosynthesis' potentially leading to future therapies centred on the more effective targeting of cancerous cells or microorganisms."
The scientists believe that they now know all the enzymes needed to produce polytheonamide from its unmodified post-ribosomal state.
Journal information: Science Express
Provided by University of Nottingham