New way to tap largest remaining treasure trove of potential new antibiotics
Scientists are reporting use of a new technology for sifting through the world's largest remaining pool of potential antibiotics to discover two new antibiotics that work against deadly resistant microbes, including the "super bugs" known as MRSA. Their report appears in the Journal of the American Chemical Society.
Sean Brady and colleagues explain that an urgent need exists for new medications to cope with microbes that shrug off the most powerful traditional antibiotics. Methicillin-resistant Staphylococcus aureus (MRSA) infections, for instance, are resistant to most known antibiotics. MRSA strikes at least 280,000 people in the U.S. alone every year, and almost 20,000 of those patients die. The typical way of discovering new antibiotics involves identifying and growing new bacteria from soil and other environmental samples in culture dishes in the laboratory. That environmental treasure-trove is the largest remaining potential source of new antibiotics. Researchers then analyze the bacteria to see if they make substances that could be used as antibiotics to kill other microbes. But most bacteria found in nature can't grow in the laboratory. That's why Brady and colleagues took a new approach to this problem.
The researchers removed DNA from soil bacteria that wouldn't grow in the lab. Then, they put this DNA into different bacteria that do grow well in culture dishes, and these bacteria acted like incubators for the new DNA. The approach enabled Brady's team to study the substances made by the soil bacteria's DNA in the lab. With this "metagenomics" method, they identified two new possible antibiotics called fasamycin A and fasamycin B that killed MRSA and vancomycin-resistant Enterococcus faecalis, which also is becoming more resistant to known antibiotics. They also determined how the new antibiotics work. "Metagenomics has the potential to access large numbers of previously inaccessible natural antibiotics," say the researchers.
More information: Environmental DNA-Encoded Antibiotics Fasamycins A and B Inhibit FabF in Type II Fatty Acid Biosynthesis, J. Am. Chem. Soc., 2012, 134 (6), pp 2981–2987. DOI: 10.1021/ja207662w
Abstract
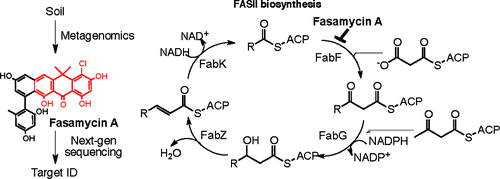
In a recent study of polyketide biosynthetic gene clusters cloned directly from soil, we isolated two antibiotics, fasamycins A and B, which showed activity against methicillin-resistant Staphylococcus aureus and vancomycin-resistant Enterococcus faecalis. To identify the target of the fasamycins, mutants with elevated fasamycin A minimum inhibitory concentrations were selected from a wild-type culture of E. faecalis OG1RF. Next-generation sequencing of these mutants, in conjunction with in vitro biochemical assays, showed that the fasamycins inhibit FabF of type II fatty acid biosynthesis (FASII). Candidate gene overexpression studies also showed that fasamycin resistance is conferred by fabF overexpression. On the basis of comparisons with known FASII inhibitors and in silico docking studies, the chloro-gem-dimethyl-anthracenone substructure seen in the fasamycins is predicted to represent a naturally occurring FabF-specific antibiotic pharmacophore. Optimization of this pharmacophore should yield FabF-specific antibiotics with increased potencies and differing spectra of activity. This study demonstrates that culture-independent antibiotic discovery methods have the potential to provide access to novel metabolites with modes of action that differ from those of antibiotics currently in clinical use.
Journal information: Journal of the American Chemical Society
Provided by American Chemical Society








.jpg)