Simulating amyloid formation

Many neurodegenerative diseases are characterized by proteins that assume an abnormal configuration, which leads to their aggregation and deposition in and around rve cells, causing cell death. This process, called amyloid formation, is a common pathological feature in diseases such as Alzheimer’s, Parkinson’s and prion diseases, as well as type II diabetes. Charlotte Hauser and co-workers from the A*STAR Institute of Bioengineering and Nanotechnology and Institute of High Performance Computing along with colleagues in Europe have now designed a class of ultrasmall peptides that simulate the self-assembly of abnormally folded proteins in such neurodegenerative conditions.
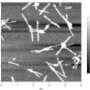
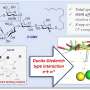
Hauser and her co-workers designed ultrasmall peptides consisting of three to six amino acid residues, each containing a characteristic motif—a ‘tail’ of uncharged residues with decreasing affinity to water capped by a polar ‘head’ residue. These peptides spontaneously self-assembled in water to form fibril structures (pictured) resembling those that make up the amyloid-β plaques found in the brains of Alzheimer’s patients.
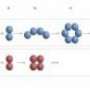
The researchers hypothesize that fiber assembly is a complex stepwise mechanism involving at least three distinct stages. Individual peptide molecules first bond to each other in an anti-parallel arrangement to form dimers. The pairs then line up to form single α-helical fibers as intermediate structures, which continue to assemble and then condense into fibrous scaffolds in the form of solid hydrogels.
The team further examined the driving forces for self-assembly and found that gel formation was critically dependent on the length of the tail and the polar nature of the head. Peptides containing six amino acid residues formed gels more readily than the others, and the strongest gels were formed by peptides containing an acidic head residue. A minimum peptide concentration was required for fiber formation, and increasing the temperature was found to accelerate the self-assembly process.
Investigation into the assembly process and experimental results were verified by computer simulations. This helped the research team confirm that the formation of peptide pairs precedes fiber formation, suggesting that the peptides have a strong tendency to aggregate because the sheet-like structures have a lower free energy state than individual fibers, and are therefore more stable.
“Understanding the driving forces that enable these ultrasmall peptides to stably self-assemble into macromolecular structures will shed light on aggregate formation in amyloidogenesis,” says Hauser. “This would facilitate the design of new therapeutics to prevent and control plaque formation in neurodegenerative disorders and a wide range of other debilitating diseases.”
More information: Hauser, C. A. E. et al. Natural tri- to hexapeptides self-assemble in water to amyloid β-type fiber aggregates by unexpected α-helical intermediate structures. Proceedings of the National Academy of Sciences 108, 1361–1366 (2011). dx.doi.org/10.1073/pnas.1014796108
Abstract
Many fatal neurodegenerative diseases such as Alzheimer’s, Parkinson, the prion-related diseases, and non-neurodegenerative disorders such as type II diabetes are characterized by abnormal amyloid fiber aggregates, suggesting a common mechanism of pathogenesis. We have discovered that a class of systematically designed natural tri- to hexapeptides with a characteristic sequential motif can simulate the process of fiber assembly and further condensation to amyloid fibrils, probably via unexpected dimeric α-helical intermediate structures. The characteristic sequence motif of the novel peptide class consists of an aliphatic amino acid tail of decreasing hydrophobicity capped by a polar head. To our knowledge, the investigated aliphatic tripeptides are the shortest ever reported naturally occurring amino acid sequence that can adopt α-helical structure and promote amyloid formation. We propose the stepwise assembly process to be associated with characteristic conformational changes from random coil to α-helical intermediates terminating in cross-β peptide structures. Circular dichroism and X-ray fiber diffraction analyses confirmed the concentration-dependent conformational changes of the peptides in water. Molecular dynamics simulating peptide behavior in water revealed monomer antiparallel pairing to dimer structures by complementary structural alignment that further aggregated and stably condensed into coiled fibers. The ultrasmall size and the dynamic facile assembly process make this novel peptide class an excellent model system for studying the mechanism of amyloidogenesis, its evolution and pathogenicity. The ability to modify the properties of the assembled structures under defined conditions will shed light on strategies to manipulate the pathogenic amyloid aggregates in order to prevent or control aggregate formation.
Provided by Agency for Science, Technology and Research (A*STAR)