With chemical modification, stable RNA nanoparticles go 3-D

(PhysOrg.com) -- For years, RNA has seemed an elusive tool in nanotechnology research -- easily manipulated into a variety of structures, yet susceptible to quick destruction when confronted with a commonly found enzyme.
"The enzyme RNase cuts RNA randomly into small pieces, very efficiently and within minutes,” explains Peixuan Guo, PhD, Dane and Mary Louise Miller Endowed Chair and professor of biomedical engineering at the University of Cincinnati (UC). "Moreover, RNase is present everywhere, making the preparation of RNA in a lab extremely difficult.”
But by replacing a chemical group in the macromolecule, Guo says he and fellow researchers have found a way to bypass RNase and create stable three-dimensional configurations of RNA, greatly expanding the possibilities for RNA in nanotechnology (the engineering of functional systems at the molecular scale).
Their results, "Fabrication of Stable and RNase-Resistant RNA Nanoparticles Active in Gearing the Nanomotors for Viral DNA Packaging,” are published online in the journal ACS Nano.
In their work, Guo and his colleagues focused on the ribose rings that, together with alternating phosphate groups, form the backbone of RNA. By changing one section of the ribose ring, Guo and his team altered the structure of the molecule, making it unable to bind with RNase and able to resist degradation.
"RNase interaction with RNA requires a match of structural conformation,” says Guo. "When RNA conformation has changed, the RNase cannot recognize RNA and the binding becomes an issue.”
While he says previous researchers have shown this alteration makes RNA stable in a double helix, they did not study its potential to affect the folding of RNA into a three-dimensional structure necessary for nanotechnology.
After creating the RNA nanoparticle, Guo and his colleagues successfully used it to power the DNA packaging nanomotor of bacteriophage phi29, a virus that infects bacteria.
"We found that the modified RNA can fold into its 3-D structure appropriately, and can carry out its biological functions after modification,” says Guo. "Our results demonstrate that it is practical to produce RNase-resistant, biologically active, and stable RNA for application in nanotechnology.”
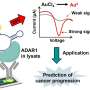
Because stable RNA molecules can be used to assemble a variety of nanostructures, Guo says they are an ideal tool to deliver targeted therapies to cancerous or viral-infected cells:
"RNA nanoparticles can be fabricated with a level of simplicity characteristic of DNA while possessing versatile structure and catalytic function similar to that of proteins. With this RNA modification, hopefully we can open new avenues of study in RNA nanotechnology.”
More information: "Fabrication of Stable and RNase-Resistant RNA Nanoparticles Active in Gearing the Nanomotors for Viral DNA Packaging", ACS Nano. pubs.acs.org/doi/abs/10.1021/nn1024658
Provided by University of Cincinnati