NIST team advances in translating language of nanopores
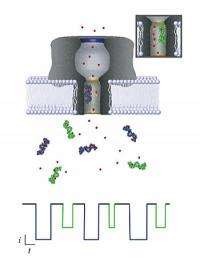
National Institute of Standards and Technology scientists have moved a step closer to developing the means for a rapid diagnostic blood test that can scan for thousands of disease markers and other chemical indicators of health. The team reports it has learned how to decode the electrical signals generated by a nanopore -- a "gate" less than 2 nanometers wide in an artificial cell membrane.
Nanopores are not new themselves; for more than a decade, scientists have sought to use a nanopore-based electrical detector to characterize single-stranded DNA for genetic sequencing applications. More recently, NIST scientists turned their attention to using nanopores to identify, quantify and characterize each of the more than 20,000 proteins the body produces—a capability that would provide a snapshot of a patient's overall health at a given moment. But while nanopores permit molecules to enter into them one at a time, determining what specific individual molecule has just passed through has not been easy.
To address this problem, members of the NIST team that previously developed a method to distinguish both the size and concentration of each type of molecule the nanopore admits have now answered the question of just how these single molecules interact with the nanopore. Their new theoretical model describes the physics and chemistry of how the nanopore, in effect, parses a molecule, an understanding that will advance the use of nanopores in the medical field.
"This work brings us one step closer to realizing these nanopores as a powerful diagnostic tool for medical science," says Joseph Reiner, who performed the work with Joseph Robertson, and John Kasianowicz, all of NIST's Semiconductor Electronics Division. "It adds to the 'Rosetta Stone' that will allow us to read what molecules have just passed through a nanopore."
Using their new methods, the team was able to model the interaction of a particular type of large molecule through a nanopore's opening with great accuracy. The molecules were polyethylene glycol (PEG), a well-understood polymer that forms chains of varying length.
"PEG chains can be very long, but each link is very small," Kasianowicz says. "It was a good test because we wanted to see if the nanopore could differentiate between two nearly identical large molecules that differ in length by only a few atoms."
The team's device was able to distinguish among different-sized PEG chains easily, and the model they have developed to describe the PEG-nanopore interactions is encouraging them to think that with further effort, the minuscule sensors can be customized to measure many different molecules quickly. "We could conceivably build an array of many nanopores, each one created to measure a specific substance," Kasianowicz says. "Because each nanopore is so small, an array with one for every protein in the body would still be tiny."
More information: J.E. Reiner, J.J. Kasianowicz, B.J. Nablo, and J.W. F. Robertson. Theory for polymer analysis using nanopore-based single-molecule mass spectrometry. Proceedings of the National Academy of Sciences, Published online on June 21, 2010, doi: 10.1073/pnas.1002194107
Provided by National Institute of Standards and Technology