DNA testing to help save corals

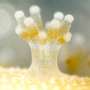
"We are seriously tackling conservation of coral reefs in Okinawa," said Dr. Chuya Shinzato of the OIST Marine Genomics Unit. Coral reefs face various extinction risks. To avert coral demise, Shinzato is providing his expertise to the Coral Reef Preservation and Restoration Project spearheaded by the Okinawa Prefectural Government. In a paper published in Frontiers in Marine Science, Shinzato and colleagues reported the establishment of DNA markers that might be applicable to all species of the Acropora reef-building coral, giving accurate identification to individual corals. The technique, similar to DNA profiling in humans, enables scientists to study genetic diversity and connectivity among the Acropora coral populations, thus finding clues to help with the conservation of coral reef ecosystems in waters around Okinawa and the world.
Coral reefs occupy only one percent of the ocean floor, but they are home to as much as a quarter of all described marine species in the world. Despite their importance, corals face a range of grave risks today, from bleaching triggered by increasing seawater temperatures, to sediment loads caused by terrestrial erosion from land development, to predation by crown-of-thorns starfish. In order to improve coral conservation and restoration efforts, it is important to increase diversity so that all coral in one area is not susceptible to the same destructive force. Diversity also helps to build reefs more robust to environmental changes than those composed of a few individual corals.
In this study, Shinzato and his team used the Acropora coral, the most common coral genera in the Indo-Pacific. They have 113 species that inhabit waters from the Red Sea through the Indian and Pacific Oceans to the Caribbean. There are four distinct groups of the Acropora coral, with a major evolutionary split occurring around 6.6 million years ago. The researchers first looked at the sequenced genome of Acropora digitifera andAcropora tenius, which belong to the two most distantly related groups. The team then located multiple repeated DNA sequences, called microsatellites in their respective genome. The number of repeats that appear differs between individual corals.
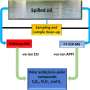
In order to detect microsatellites commonly present in the two Acroporaspecies used in the study, the researchers used a process called polymerase chain reaction (PCR), using a pair of flanking DNA sequences that extend to either side of the microsatellite region. The distance between these neighboring sequences, called primers, differs between each individual coral due to the number of times the microsatellites are repeated, providing valuable information on the relatedness of different individuals. However, the development of microsatellite DNA markers remains time consuming, expensive, and labor intensive since the markers are often species specific. Despite these drawbacks, Shinzato and colleagues developed 14 microsatellite DNA markers found in two of the most diverse Acroporaspecies. Since the 14 markers have been conserved during their evolution dating back to 6.6 million years ago, the researchers assert that these markers should also be present in all of the other 111 species of Acropora corals. The study spares scientists the time and labor of creating markers for the other 111 species and serves as a powerful research tool to identify individual corals, thus contributing to population genetics studies and conservation of Acropora corals.
The work has direct implications for Okinawa. Since the Acropora coral is the most common coral in the area, the study can be applied to most of Okinawa's coral reefs. Already, the Okinawa Prefectural Government has taken steps to preserve and regenerate corals in nearby waters by employing this technology to increase genetic diversity in coral plantation.
"I truly hope the technique we have developed will contribute to coral reef transplantation and restoration," said Shinzato. The markers established by the OIST team can be applied to all Acropora corals in the world. The study may pave a way for global restoration of the rich coral reef ecosystems that are suffering.
More information: "Development of novel, cross-species microsatellite markers for Acropora corals using next-generation sequencing technology." Chuya Shinzato, et al. Front. Mar. Sci., 23 May 2014 | DOI: 10.3389/fmars.2014.00011
Provided by Okinawa Institute of Science and Technology