Research may unlock enzyme's role in disease
A UT Arlington chemist doing National Science Foundation-funded research on enzymes that regulate human biology has uncovered characteristics that could be used to identify predisposition to conditions such as heart disease, diabetic ulcers and some types of cancer.
Brad Pierce, an assistant professor of chemistry/biochemistry at The University of Texas at Arlington, recently led a team that examined an oxygen utilizing iron enzyme called cysteine dioxygenase or CDO, which is found in high levels within heart, liver, and brain tissues. Enzymes are proteins that act as catalysts to enable metabolic functions, but under some circumstances these oxygen-dependent enzymes can also produce highly toxic side products called reactive oxygen species or ROS.
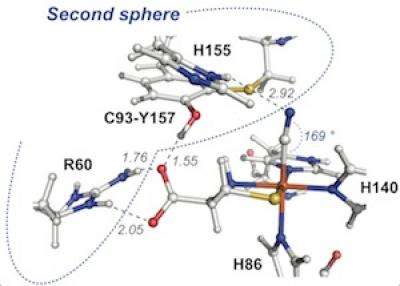
For the first time, Pierce's team found that mutations outside the CDO active site environment or "outer coordination sphere" have a profound influence on the release of ROS. Excess ROS has been linked to numerous age-onset human disease states.
"Most research in the past has focused on the active site inner coordination sphere of these enzymes, where the metal molecule is located," said Pierce. "What we're finding is that it's really the second sphere that regulates the efficiency of the enzyme. In essence, these interactions hold everything together during catalysis. When this process breaks down, the enzyme ends up spitting out high levels of ROS and increasing the likelihood of disease."
The study was published in December by the American Chemical Society journal Biochemistry. Pierce is corresponding author on the paper, with UT Arlington students Wei Li, Michael D. Pecore and Joshua K. Crowell as co-authors. Co-author Elizabeth J. Blaesi is a graduate research assistant at the University of Wisconsin.
Pierce believes the findings from the CDO enzyme could be applied to other oxygen-dependent enzymes, which make up about 20 percent of the enzymes in the human body.
"In principle, these findings could be extended to better understand how other enzymes within the class generate ROS and potentially be used to screen for genetic dispositions for ROS-related diseases," he said.
Pierce's research brings a new level of detail to enzyme study through the use of electron paramagnetic resonance or EPR, a technology similar to the magnetic resonance imaging or MRI used in the medical field. In fall 2012, the National Science Foundation awarded Pierce a three-year, $300,000 grant to study enzymes that are catalysts for the oxidation of sulfur-bearing molecules in the body.
"Dr. Pierce's research is a good example of how basic science can set a path toward discoveries that affect human health. We look forward to his continued exploration of these findings," said Pamela Jansma, dean of the UT Arlington College of Science.
More information: Biochemistry. 2013 Dec 23;52(51):9104-19. DOI: 10.1021/bi4010232.
Journal information: Biochemistry
Provided by University of Texas at Arlington