Newly discovered protist suggests evolutionary answers, questions
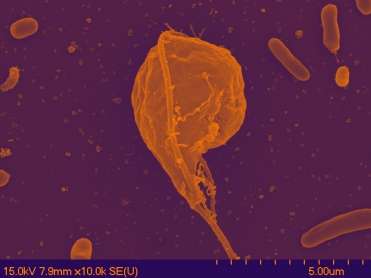
From Massachusetts to Mississippi, a unicellular protist is hinting at answers about the evolution of multicellularity while raising a whole new set of questions.
Matthew Brown, assistant professor of biological sciences at Mississippi State University, recently led a research team that identified the protist as a new organism and classified its genomics.
Jeffrey Silberman collected sediment specimens in Marstons Mills, a village in Barnstable, Mass., and the University of Arkansas associate professor isolated an organism he found. Since Brown had begun post-doctoral work in genomics at Dalhousie University in Nova Scotia, Silberman offered his former UA doctoral student the opportunity to name and classify it on the evolutionary tree of life.
Brown headed the investigation that discovered the unicellular organism's proteins and genes are similar to those found in multicellular life-forms. The protist Pygsuia biforma belongs to a newly identified group they named "Obazoa," which is closely related to animals and fungi.
"We then looked for specific multicellular toolkit genes, and we found genes that scientists had believed to be animal-specific," Brown said. "Integrins and the whole suite of proteins that work with integrins were thought to be something innate to multicellularity and used only for cell-to-cell communication.
"This discovery shows that these genes have been co-opted for a different use. We don't know what it does in unicellular organisms, but we can now place the origin of genes that are associated with multicellularity in unicellular organisms."
Additionally, the anaerobic protist has mitochondria, energy factories that produce adenosine triphosphate, or ATP. Brown said ATP production typically requires oxygen, but the protist lives in oxygen depleted environments. As a result, Pygsuia biforma raises questions related to the presence and function of mitochondria in anaerobic unicellular organisms.
These discoveries and new research questions they raise are important because they offer new insights into the science of evolution, Brown explained.
"By tracking the evolutionary history of these particular organisms, we're able to look at ancestral states of certain gene suites, and that's the really important thing—we need a better understanding of protist diversity and protist genome evolution to understand how organisms like animals evolved," Brown said.
Evidently, the international scientific community agrees: The team's research paper detailing these discoveries, "Phylogenomics demonstrates that breviate flagellates are related to opisthokonts and apusomonads," was recently published in Proceedings of the Royal Society B, the leading United Kingdom biological research journal.
Because of Brown's bioinformatics expertise in genetic and protein sequencing, as well as his leadership role in documenting the protist's morphology, he was the paper's lead author.
His work continues in the MSU biological sciences' Evolutionary Protistology Laboratory, also known on campus as Brown's Lab. Work there examines the evolution of eukaryotic lineages with comparative genomics and developmental transcriptomics.
Journal information: Proceedings of the Royal Society B
Provided by Mississippi State University















