Temperature-dependent radiolysis reveals dynamics of bound protein waters
(Phys.org) —Water is crucial to the functioning of the body, even on very small scales. The ubiquitous liquid is key to the structure, folding and stability of proteins, but one of the still unanswered questions in the study of the structure and function of proteins and DNA is their exact relationship to their water environment. All of the molecules in our bodies function in water, but until now, we haven't had a lot of experimental techniques to understand what water is doing or where it is binding to the interior surfaces of proteins.
A team of scientists from Case Western Reserve University used the National Synchrotron Light Source (NSLS) at Brookhaven National Laboratory to develop a technique that pinpoints the location and motion of water molecules bound to proteins. Using temperature-dependent radiolysis and mass spectroscopy, they are able to identify where water is binding tightly or loosely on the surface of a protein and how it is influencing a protein's function.
"It's as if there were a window ledge with a pebble stuck in it, so the window doesn't shut tightly," said Mark Chance, director of the Center for Proteomics and Bioinformatics at Case Western University. "The water is like that pebble. It could be an obstacle to the formation of the protein complex, just like the pebble stops the window closing. Or, it could be in just the right position, as if the window had a small notch carved in just the shape of the pebble. Both of those situations could occur in nature."
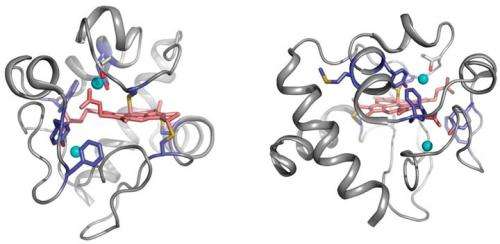
With so much water in a protein's environment, it's difficult to pick out and study individual molecules because they have such small signals within the sea of water they inhabit. So, in order to find water that's tightly bound to a protein, the team chemically activated protein waters with an x-ray beam, which permanently attached the tightly bound waters to the protein. They then wash out the system and take a measurement of the mass of the protein. A shift in mass tells them where the water is bound tightly.
Measuring the exchange rate of water (16O) versus heavy water (18O) shows how fast the mass of the peptide changes due to bound water molecules. "To our surprise, that process was slower than we thought it would be," Chance said. "The water is more tightly bound than we would have expected, and on a timescale that's important for the chemistry of protein reactions."
Chance said this method could give insight into the function of water in enzymes such as hexokinase, which catalyzes the first step in oxidization of food into energy in the body. Hexokinase takes glucose from food, puts a phosphate on it and traps it. "We know it works by using a lid with a cap that lets the molecule in and then the lid flips down. That lid has waters decorating the surface and now we can understand things about it for the first time," he said. "How tightly bound are the waters on that lid? And how does the structure of the water manipulate the time scale of the lid and manipulate energy production in the body? Those are the questions this technique will allow us to answer."
The method is also well suited to studying aquaporins – proteins embedded in cell membranes that regulate water flow – or ion channels, which transport ions along with water. "This is one of the most interesting things I've done in a number of years," Chance said. "People are really excited about the possibilities this opens up to study what water is doing and where it is. And this opportunity to study the structure of water has a bright future at the National Synchrotron Light Source II." Now under construction at Brookhaven Lab, that new facility will begin operating in 2015.
Chance was recently awarded the largest instrumentation grant from the National Science Foundation for a state-of-the-art wiggler to be used at the beamline he is overseeing at NSLS-II. The instrument will allow scientist to probe the structures of molecules in vivo and to answer fundamental questions about structural biology.
More information: www.bnl.gov/today/docs/PDF/PNA … teinWaters-Slide.pdf
Provided by Brookhaven National Laboratory