The cell that isn't: New technique captures division of membrane-less cells
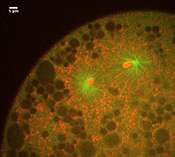
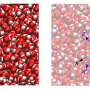
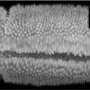
This may look like yet another video of a dividing cell, but there's a catch. You are looking at chromosomes (red) being pulled apart by the mitotic spindle (green), but it's not a cell, because there's no cell membrane. Like a child sucking an egg out of its shell, Ivo Telley from the European Molecular Biology Laboratory (EMBL) in Heidelberg, Germany, removed these cellular 'innards' from a fruit fly embryo, at a stage when it is essentially a sac full of membrane-less 'cells' that divide and divide without building physical barriers to separate themselves from each other.
"It's the first time we can study ongoing cell division without the cell membrane, and that means we can physically manipulate things," says Telley, "so we can uncover the physical forces involved, and see what are the constraints."
The new technique is described in detail today in Nature Protocols, and has already led Telley and colleagues to a surprising discovery. They found that, although successive divisions fill the embryo with more and more material, leaving less and less space for each spindle, and spindles become smaller as the embryo develops, simply squeezing the 'cell' into tighter quarters doesn't make it produce a smaller spindle.
Combined with the genetic manipulation approaches commonly used in fruit fly studies, the scientists believe their new technique will help to unravel this and other mysteries of how a cell becomes two.
The work, which started in Thomas Surrey's lab at EMBL, was carried out by Telley and Imre Gáspár in Anne Ephrussi's lab at EMBL. Surrey is now at Cancer Research UK.
More information: A single Drosophila embryo extract for the study of mitosis ex vivo. Telley, I.A., Gáspár, I., Ephrussi, A. & Surrey, T. Nature Protocols, Advanced online publication on 17 January 2013. DOI: 10.1038/nprot.2013.003
Abstract
Spindle assembly and chromosome segregation rely on a complex interplay of biochemical and mechanical processes. Analysis of this interplay requires precise manipulation of endogenous cellular components and high-resolution visualization. Here we provide a protocol for generating an extract from individual Drosophila syncytial embryos that supports repeated mitotic nuclear divisions with native characteristics. In contrast to the large-scale, metaphase-arrested Xenopus egg extract system, this assay enables the serial generation of extracts from single embryos of a genetically tractable organism, and each extract contains dozens of autonomously dividing nuclei that can be prepared and imaged within 60–90 min after embryo collection. We describe the microscopy setup and micropipette production that facilitate single-embryo manipulation, the preparation of embryos and the steps for making functional extracts that allow time-lapse microscopy of mitotic divisions ex vivo. The assay enables a unique combination of genetic, biochemical, optical and mechanical manipulations of the mitotic machinery.
Provided by European Molecular Biology Laboratory