As the worm turns, its secrets are revealed
An international team of scientists, led by researchers at the University of California, San Diego School of Medicine, have developed a new method for discerning the functions of previously uncharacterized genes and placing them in interactive, functional networks that reveal how gene products interact to bring about cellular events.
The research is published in the April 29 issue of the journal Cell. It was led by principal investigators Karen Oegema, PhD, professor of cellular and molecular medicine and head of the Laboratory of Mitotic Mechanisms in the Ludwig Institute for Cancer Research at UC San Diego, and Kristin C. Gunsalus, PhD, assistant professor in the Center for Genomics and Systems Biology in the Department of Biology at New York University.
More than a decade of genome sequencing projects have generated a comprehensive "parts" list of the genes required to build an organism, an inventory of the necessary cellular building blocks. But the functions of many of these genes remain unknown, preventing researchers from fully deciphering their cellular pathways and how their interactions might shed light on human disease.
One of the stars of this research is Caenorhabditis elegans, a tiny, much-studied worm that is an important model system for understanding processes in animal cells. In recent years, scientists have sought to create systemic catalogs of its gene functions, and those of other model organisms. These large-scale efforts place genes in interactive networks. Within these networks, proximity reflects similarity of function. In other words, genes with similar functions are directly linked and genes with dissimilar functions are further apart. The functions of uncharacterized genes are inferred based upon their proximity to genes whose functions are known.
Generating functional maps with the power to resolve differences in gene function requires a lot of information, which is typically obtained by "high-content" screening in which genes are individually inhibited and the consequences are documented by filming the behavior of individual cells.
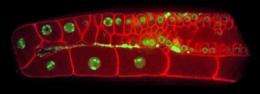
Oegema, with the study's first author, Rebecca Green, PhD, a postdoctoral fellow at the Ludwig Institute and UCSD School of Medicine, decided to take a different approach. "Rather than monitoring individual cells, we monitored the effect of gene inhibitions on the structure of a complex tissue in a multicellular organism," said Green. "In this case, the reproductive organ of C. elegans."
The scientists discovered that inhibiting different genes in the nematode produced a remarkably diverse and information-rich spectrum of effects on tissue structure, essentially creating a "fingerprint" for each gene that allowed them to predict its function.
Oegema and colleagues also developed a new method to quantitatively assess the significance of gene-gene connections, allowing them to translate the information into a functional gene network. In collaboration with Gunsalus and colleagues, who developed a Java-based tool for visualizing gene networks, they produced an integrated functional network for a set of 818 essential C. elegans genes, which has been made available to interested researchers.
The network, the scientists said, will be useful for predicting the function of related human genes and the broader approach may be useful for generating functional gene networks in other organisms, including vertebrates.
Provided by University of California - San Diego















