Physicists unravel mystery of repetitive DNA segments

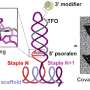
With new tools that can grab individual strands of DNA and stretch them like rubber bands, Rice University scientists are working to unravel a mystery of modern genomics. Their latest findings, which appear in Physical Review Letters, offer new clues about the physical makeup of odd segments of DNA that have just one DNA base, adenine, repeated dozens of times in a row.
These mysterious "poly(dA) repeats" are sprinkled throughout the human genome. Scientists have also found them in the genomes of animals, plants and other species over the past decade. But researchers do not know why they are there, what function they perform or why they occur only with the DNA base adenine and not the other three DNA bases -- cytosine, guanine and thymine.
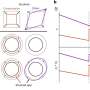
"Previous investigations of poly(dA) have suggested that adenine bases stack in a very uniform way," said Ching-Hwa Kiang, a co-author of the new study and assistant professor of physics and astronomy at Rice. "Our investigation focused on what happens when single strands of poly(dA) were stretched and these stacks were pulled apart."
Kiang's research group specializes in studying the physical and mechanical properties of proteins and nucleic acids, and their primary tool is one of the mainstays of nanotechnology research -- the atomic force microscope, or AFM. The business end of an AFM is like a tiny phonograph needle. The tip of the needle is no more than a few atoms wide, and the needle is at the end of an arm that bobs up and down over the surface of what is being measured. While nanotechnologists use the device to measure the thickness of samples, Kiang's group uses it in a different way.
To begin her experiments, Kiang first places a thin coating of the proteins she wishes to study on a flat surface. This is placed under the AFM arm so the bobbing AFM needle can dip down and grab the ends of one of the proteins. As the arm retracts, it unravels the protein.
All proteins fold into a characteristic shape. Like tiny springs, they remain in this compact "lowest energy" state unless they are pried apart.
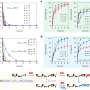
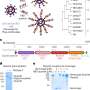
The new study on poly(dA) was conducted by Kiang, Rice graduate student Wuen-shiu Chen and colleagues at Rice and National Chung Hsing University (NCHU) in Taiwan. The team discovered that poly(dA) behaves differently depending upon the speed with which it is stretched. When the AFM bobbed rapidly, the poly(dA) segments behaved like any other segment of single-stranded DNA. But when the AFM motion was slowed, the team found that the amount of force required to stretch the poly(dA) changed. At two particular locations, the strand lengthened for a short distance without any additional force at all.
"Typically, single strands of DNA behave like a rubber band: The resistance increases as they stretch, meaning you have to pull harder and harder to continue stretching them," Kiang said. "With poly(dA), we found these two points where that doesn't apply. It's as if you have to pull harder and harder, and then for a brief time, the band stretches with no additional force whatsoever."
Kiang said the exact causes and implications of the phenomenon are unclear. But scientists know that double-stranded DNA must be pried apart at discrete locations so that the cell's machinery can read the genetic code and convert it into proteins. There has been some speculation that the adenine repeats play a role in ordering genomic information; Kiang said the new findings raise even more questions about the role the repeats might play in gene regulation and genome packaging and how they might be potential targets for cancer drugs.
Provided by Rice University