Cornell researchers reveal structure of key protein
(PhysOrg.com) -- For the first time, researchers -- all Cornell scientists -- have characterized the structure of a protein that belongs to certain enzymes that are essential for proper functioning in all life forms, from yeast to humans.
The enzymes, belonging to the so-called Sac family, are involved in cellular signaling and membrane trafficking. Scientists have found that when the gene that expresses Sac enzymes is deleted in animals, the animals die, and mutations of related genes in humans lead to cancers and such neurodegenerative hereditary diseases as Charcot-Marie-Tooth Type 4J (CMT4J) and Lou Gehrig's disease.
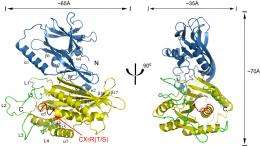
Researchers from Cornell's Weill Institute of Cell and Molecular Biology, reporting online in the Journal of the European Molecular Biology Organization, have characterized for the first time the crystal structure of the Sac1 protein in yeast. Yeast serves as a model organism for all cells; most of the 6,000 genes in yeast are also found in humans. The Sac1 protein in yeast is a progenitor for related Sac proteins also found in plants and animals.
Understanding the Sac1 protein's structure opens the way for experiments that may reveal how these fundamental enzymes interact with cell membranes to enable essential cellular processes, which could also lead to drugs that target related diseases.
"This enzyme was first discovered in 1989, but no one had seen the atomic structure of this protein," said Yuxin Mao, an assistant professor of molecular biology and genetics and the paper's senior author. Andrew Manford, a graduate student in the lab of Scott Emr, director of the Weill Institute, is the paper's lead author. "Others have tried, but this is the first time" the protein's structure has been revealed, said Mao.
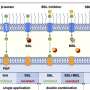
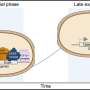
Much like an on and off switch, pathways that signal cells to divide, migrate or transport materials in and out of the cell are often activated by attaching a phosphate group to proteins or lipids (a process called phosphorylation) and similarly deactivated by the removal of the phosphate group. A class of enzymes called phosphatases mediates the removal of phosphates, and the Sac family of enzymes the Cornell researchers studied are lipid phosphatases. Such diseases as CMT4J and Lou Gehrig's disease occur when Sac family phosphatases fail to function properly, leading to a buildup of a group of phosphorylated lipids.
Mao and colleagues determined the structure by growing Sac protein crystals, which allowed researchers to view a protein's atomic structure through X-ray diffraction. Mao's lab used Cornell's synchrotron to solve the crystal structure of the Sac1 protein at an atomic resolution of less than 2 angstroms (two ten-millionths of a millimeter).
"This opens up biochemical studies of how these enzymes function -- it's a breakthrough in this direction of study," said Mao. "And it helps our studies of other members of the Sac family."
Provided by Cornell University