Stable Thanks to Dynamics - DNA Component Resists UV Radiation
(PhysOrg.com) -- Complex computer simulations have, for the first time, allowed scientists to examine in detail the processes that help to ensure the stability of DNA when exposed to UV light. The findings, achieved primarily in relation to DNA component 9H-adenine, have been published in the Journal of the American Chemical Society (JACS). Moreover, in recognition of the high quality of the work, they have also been posted in the publication's newly established online section JACS Select. The results of the project, which was supported by the Austrian Science Fund FWF, show that an ultrafast, two-step process forms one basis for the photostability of DNA.
UV rays don't just tan our skin - they can also "rob" atoms of their electrons, thereby destroying organic compounds. In order to counteract these destructive effects, the DNA component adenine uses an ultrafast process that lasts for less than one trillionth of a second (a picosecond). UV light causes excitation of the electrons in adenine, which in turn causes the electrons initially to reach a higher energy state. They then return to their original state - this happens at breathtaking speed in the case of adenine. During this process, potentially damaging excitation energy is converted into harmless vibrational energy within the molecular skeleton. This protects the DNA against damage. While scientists were previously aware of this process, they had not been able to simulate it due to the extreme complexity involved and had therefore been unable to study it in detail - until now.
Simulating a Transition
The innovative use of a computational method aimed at simulating the dynamics of quantum states enabled the Vienna-based team to make accurate statements about the mechanisms on which the photostability of adenine is based. Prof. Hans Lischka, who together with Prof. Mario Barbatti is leading the team at the Institute of Theoretical Chemistry of the University of Vienna explains: "Due to the size of the molecule, the relatively long simulation times, and the complexity of the electronic spectra, this project was a formidable challenge. And taking on this challenge has paid off".
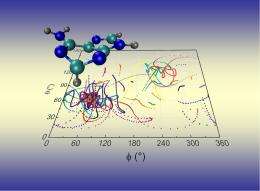
Lischka's team calculated in detail the transition between the individual energy states of the electrons coupled with the movement of the atomic nuclei. Initial data show that this transition is not uniform, but rather a process made up of two steps. The first of these is "ultrashort" and lasts for only 22 femtoseconds (22 quadrillionths of a second). During this first step, the electrons drop from a high energy state (S3) to a lower state (S1). The second step lasts around 20 times longer than the first one, or half a picosecond. After this step, the adenine electrons will have returned to their original low-energy state (S0) - at unimaginable speed.
Motion of Electrons and Atomic Nuclei
Commenting on the results, the highly regarded American Chemical Society highlighted a number of points in the project, including the number of nuclear paths calculated by Lischka and his team. During their work, the team simulated not just one or two, but 60 of these paths, which are also referred to as trajectories. Simulating this many trajectories, enabled the team to compile a statistics for each process responsible for the photostability of adenine. Lischka and Barbatti on their approach: "By using a multitude of trajectories, we were able to draw statistical conclusions. For example, we found that in 98% or practically all of the trajectories we calculated, the S1 state was reached 60 femtoseconds after exposure to UV light". The computer simulation method used by Lischka and Barbatti in this specific context holds a great deal of potential for further research on ultrafast photochemical processes in polyatomic molecules. During the current project, the potential of this exceptionally time-intensive method was utilized in full, thanks to the support that Lischka's work received from the Austrian Science Fund FWF. The resulting insights and the inclusion of Lischka and Barbatti's publication in the JACS Select online collection of innovative developments clearly demonstrate that Lischka has mastered his "formidable challenge".
More information: "Nonadiabatic Deactivation of 9H-Adenine: A Comprehensive Picture Based on Mixed Quantum-Classical Dynamics" M. Barbatti & H. Lischka. J. Am. Chem. So., 2008, 130 (21), 6831-6839, DOI:10.1021/ja800589p
JACS Select: pubs.acs.org/JACSbeta/jvi/issue3.html
Provided by Austrian Science Fund (FWF)