This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
proofread
Fast track to food safety: New test spots seafood pathogen in 30 minutes
Vibrio parahaemolyticus is a Gram-negative, halophilic bacterium prevalent in marine environments and is the primary cause of acute hepatopancreatic necrosis, also known as early death syndrome, in aquaculture.
It represents a considerable public health hazard, especially through the consumption of raw or undercooked seafood. The bacterium can contaminate seafood surfaces, leading to foodborne outbreaks. Current detection methods, which rely on microbial isolation, culturing, and biochemical identification, are too slow for effective point-of-care testing (POCT).
In a notable advancement for food safety, scientists from the Shanghai Academy of Agricultural Sciences have unveiled a novel detection platform that identifies Vibrio parahaemolyticus within 30 minutes. This innovation could significantly reduce the risk of foodborne illness from seafood. Published in Food Quality and Safety February 2024, this method marks a substantial improvement in food safety and public health measures.
The research team developed an innovative platform that swiftly detects the presence of Vibrio parahaemolyticus in seafood. This rapid-response system is transformative for food safety, where early detection of pathogens is crucial for preventing illness.
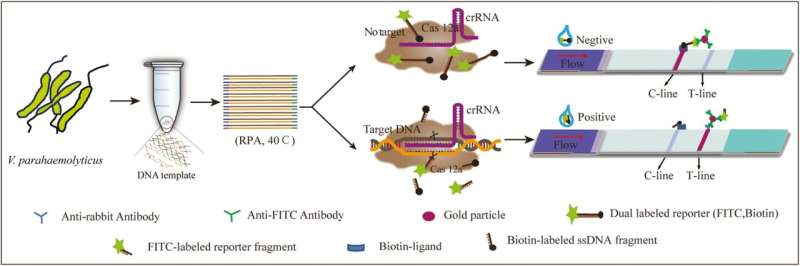
The platform utilizes a combined approach involving recombinant polymerase amplification (RPA), the CRISPR/Cas12a system, and an immunochromatographic test strip (ICS). It specifically targets the tlh gene of V. parahaemolyticus, facilitating highly sensitive detection.
The procedure starts with extracting bacterial DNA from the seafood sample, followed by RPA for amplification. The CRISPR/Cas12a system then accurately identifies and cleaves the target gene, with the ICS providing a visual confirmation of the bacterium's presence. This method achieves a detection limit of 2.5×102 fg/µL for plasmid DNA and 1.4×102 CFU/mL for the bacteria.
Remarkably, it can detect V. parahaemolyticus in salmon sashimi at concentrations as low as 154 CFU/g without sample enrichment. This breakthrough overcomes the drawbacks of traditional culture-based methods, offering a faster, more accessible approach for monitoring seafood safety.
Dr. Haijuan Zeng, the corresponding author and leader of the Biotechnology Research Institute at the Shanghai Academy of Agricultural Sciences, stated, "Our innovative detection platform represents a significant advancement in the rapid and sensitive detection of Vibrio parahaemolyticus, proving especially valuable for ensuring seafood safety and preventing public health crises."
This new method could revolutionize how food safety is monitored in the seafood industry, offering a rapid, cost-effective solution that can be implemented directly at points of sale or during food handling, significantly shortening the detection timeframe and potentially averting foodborne outbreaks before contaminated products reach consumers.
More information: Jinbin Wang et al, A visual, rapid, and sensitive detection platform for Vibrio parahaemolyticus based on RPA-CRISPR/Cas12a and an immunochromatographic test strip, Food Quality and Safety (2024). DOI: 10.1093/fqsafe/fyae008
Provided by TranSpread