February 15, 2024 feature
This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Detection of a new state in the protein folding process
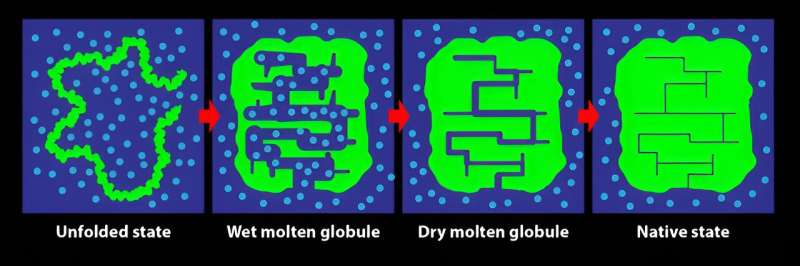
Scientists have discovered a new, intermediate state in the process of protein folding, showing folding can occur in two stages, one fast and the next found to be much slower. The findings are published in the journal Physical Review Letters.
After a protein—a chain of up to 20 different amino acid molecules—is created in a cell, it spontaneously coils up ("folds") from a random structure into an ordered three-dimensional one that makes it biologically useful. (Improper folding can result in diseases such as amyotrophic lateral sclerosis, ALS, also known as Lou Gehrig's disease.)
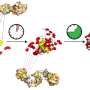
For simpler proteins, this typically occurs very quickly, over about 0.01 milliseconds (ms), into its nearly complete form. In this research, a second, slower state of 3–10 milliseconds duration was found to occur after the first step, up to a thousand times longer than the prior folding, due to small rearrangements of sidechains on the protein. (By contrast, the blink of a human eye lasts between 100 and 400 milliseconds.)
Scientists have learned to predict, using machine learning, what configuration a protein will fold into. But it's only poorly understood why each particular structure, different for each protein, is achieved, and how it happens as the various amino acids chemically react with one another.
Prior methods of analysis have, by methods such as optical spectroscopy (which measures the difference between how left- and right-handed circularly polarized light is absorbed) looked at how proteins squeeze water from within them as they turn and twist into their coiled, folded state. However, such optical measurements do not capture the final, long-term folding discovered here.
Researchers at the National Institute of Diabetes and Digestive and Kidney Diseases (NIDDK) of the National Institute of Health in the U.S, led by Robert Tycko, the Acting Chief of its Laboratory of Chemical Physics, decided to look for such longer-term folds.
They chose the protein HP35, known for its simplicity and fast folding, consisting of 35 amino acid residues. (Residues are the result of a link between two amino acid molecules, forming a peptide, which removes water. These residues influence other residues and the resulting chemical properties of the entire protein.) Also known as the "villin headpiece subdomain," it is a protein commonly used in folding studies.
The group used heat to unfold HP35, heating it over 25 ms to a hot 95°C. They then allowed it to refold during an "incubation period" of about 0.1 ms at a warm 30°C. They kept the protein at this temperature for an incubation period period of 1 to 10 milliseconds, then flash froze it to -35°C, where particles of about 0.3 millimeters in diameter form.
After observing HP35's fast folding using optical spectroscopic probes, the slurry of frozen particles was analyzed using solid state nuclear magnetic resonance of carbon-13 atoms in the protein residues to observe their behavior after the initial fast folding. At 13C's nuclear spin characteristic oscillation frequencies, the signal of the oscillating applied magnetic field becomes sharper when the molecular structure becomes more rigid.
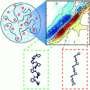
The group observed that the 13C resonance frequencies for two specific residues did not change during the incubation period, indicating they were sections of the protein that had already folded by the beginning of the incubation period. However, the signal from the resonance frequencies of several other residues in other parts of the protein sharpened over the 10-ms incubation period, revealing that those protein sections were still folding after the start of the incubation period.
That is, HP35 appeared to be folded over 0.1 ms at the beginning of the incubation period, but actually becomes fully folded only after the end of the 10-ms incubation period, becoming rigid in its final form.
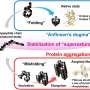
This result is the first definite evidence for the formation of what's called the DMG (dry molten globule) state during folding. The DMG was first proposed theoretically as the penultimate state taken by a folding protein in 1989—until then, only the wet molten globule state, with the pre-formed protein in a larger configuration within water, had been seen.
Until now there had been no evidence for the DMG state, but there were suggestions it existed during studies of the unfolding of various proteins. Another study of an enzyme—catalyst proteins that accelerate chemical reactions—provided hints that the folding may continue over a period of days.
"The second stage of structural optimization or annealing was not observed in previous studies but shows up in our experiments because we measure NMR signals," said HIDDK's Tycko, "which are very sensitive to conformational variations and local structural details."
Noting that much of the equipment to measure the sub-millisecond temperature jumps was developed in his lab, he added that "with this equipment, we can look for similar effects in other biomolecular processes that involve large unidirectional structural changes."
More information: C. Blake Wilson et al, Experimental Evidence for Millisecond–Timescale Structural Evolution Following the Microsecond–Timescale Folding of a Small Protein, Physical Review Letters (2024). DOI: 10.1103/PhysRevLett.132.048402
Journal information: Physical Review Letters
© 2024 Science X Network