This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
proofread
Improving grape yield predictions: The rise of semi-supervised berry counting with CDMENet
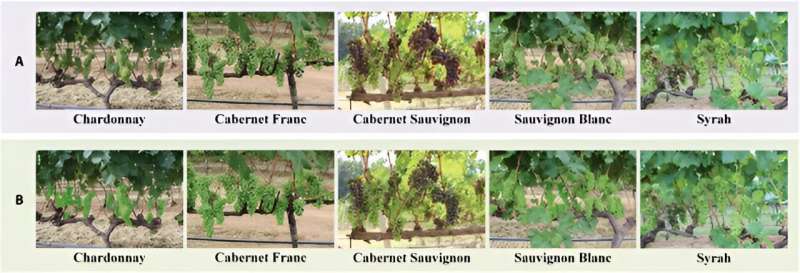

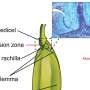
To improve grape yield predictions, automated berry counting has emerged as a crucial yet challenging task due to the dense distribution and occlusion of berries. While grape cultivation is a significant global economic activity, traditional manual counting methods are inaccurate and inefficient.
Recent research has shifted towards deep learning and computer vision, employing detection and density estimation techniques for more precise counts. However, these methods grapple with the variability of farmland and high occlusion rates, leading to significant counting errors.
Additionally, creating high-performance algorithms demands expensive data labeling, making it a significant hurdle for widespread use and adaptability in the field. The need for a cost-effective, accurate automated counting method remains a pressing research problem.
In November 2023, Plant Phenomics published a research article titled "Semi-supervised Counting of Grape Berries in the Field Based on Density Mutual Exclusion."
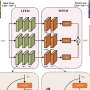
To effectively and cost-efficiently count grape berries, the research presents CDMENet, a semi-supervised method that uses VGG16 for image feature extraction and density mutual exclusion to understand spatial patterns and unlabeled data.
The method also employs a density difference loss to amplify feature differences between varying density levels. Conducted on an Ubuntu system with Python and deep learning frameworks, the algorithm's performance was tested using a specified hardware setup, preprocessing techniques, and training details such as learning rates and optimization methods.
The results were promising, with CDMENet outperformed both fully and semi-supervised counterparts in terms of Mean Absolute Error (MAE), Root Mean Square Error (RMSE), and the coefficient of determination (R2).
Particularly notable was its performance with limited labeled data, showcasing superior accuracy and reduced errors compared to other models. Additionally, the method's robustness was further evidenced in various ablation studies, demonstrating the significant roles of unlabeled data, density difference loss, and the number of auxiliary task predictors.
Despite minor influences from prediction confidence thresholds, CDMENet maintained stable and robust performance.
In conclusion, CDMENet presents a viable solution for grape berry counting in fields, reducing the need for extensive manual labeling while improving accuracy and reducing errors.
Its efficient use of unlabeled data, enhanced feature representation through density difference loss, and overall robust performance against a backdrop of varying conditions underscore its potential as a cost-effective tool in agricultural yield estimation.
Future work might explore optimizing loss functions further and deploying the algorithm in field robots or other practical applications.
More information: Yanan Li et al, Semi-supervised Counting of Grape Berries in the Field Based on Density Mutual Exclusion, Plant Phenomics (2023). DOI: 10.34133/plantphenomics.0115
Provided by Plant Phenomics