This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
New tech enables scientists to see living cells' organelles in motion at super-high resolution
The research group of Professor Yoav Shechtman from the Israel Institute of Technology Faculty of Biomedical Engineering has developed groundbreaking technology enabling scientists to see dynamic processes in living cells. Their study was published in Nature Methods.
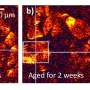
Until now, high-resolution microscopy enabled researchers to observe sub-cellular structures such as organelles, but at the cost of a long acquisition time—a minute or more per image—so whatever was being looked at needed to be held perfectly still.
This posed a real problem for biologists, since living cells, and the organelles inside them, are naturally in constant motion. One can fix them in place artificially, but then they are not in their natural state.
A study led by Ph.D. student Alon Saguy and Prof. Yoav Shechtman offers an innovative solution that uses artificial intelligence (AI) to enable scientists to see sub-cellular dynamics without being compromised by long acquisition times.
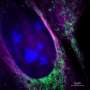
How does one find what they are looking for under a microscope? In biology, scientists commonly use fluorescent dyes to stain specific structures of interest. This creates high-contrast images of the labeled structures, which can then be seen clearly. There is, however, a physical limit on how good a resolution one can achieve using this methodology. It cannot resolve objects smaller than 200nm—approximately half the wavelength of visible light.
For some uses, a resolution of 200nm is good enough. But many structures in the cell are much smaller. Microtubules, which form the cell's "skeleton," for example, are only ~25nm thick. For the methodology to make such structures visible, Professors Eric Betzig, Stefan Hell, and William E. Moerner received the Nobel Prize in Chemistry in 2014.
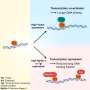
The technique Betzig developed, called single-molecule localization microscopy (SMLM) relies on making not a single image, but a video recording of the fluorescently labeled sample. In each frame, only a few individual molecules emit light, creating a sparse speck pattern. Each speck of light is localized in high resolution and the localizations of the entire video are stacked together to form a high-resolution image.
SMLM has a significant drawback: since exposures of over a minute are required to generate a single high-resolution image, the cell must be fixed, like old photos, which required subjects to stand still for a long time, lest the image come out blurry.
And just as a photo—or even more so, a video—of a child in play or an athlete mid-jump is truer to life than the postured old photographs, scientists need to see the cell and the organelles within it move, respond to stimuli, and do the things that they naturally do.
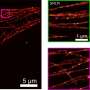
The Technion team developed a clever solution. "Things move in a living cell, but they move with a certain regularity," Prof. Shechtman explained. "If we look at microtubules, for example, they are sort of like threads, bound together into a mesh. They move, but you don't have bits of them randomly hopping about. There's a pattern to the movement."
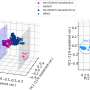
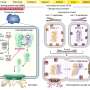
Artificial Neural Networks (ANNs) are powerful AI tools that are very good at finding patterns. Prof. Shechtman and his team trained their ANN to find patterns in SMLM videos. The ANN would receive the recording frame by frame, each frame showing only a few spots of light and produce a continuous video of the structures behind those spots.
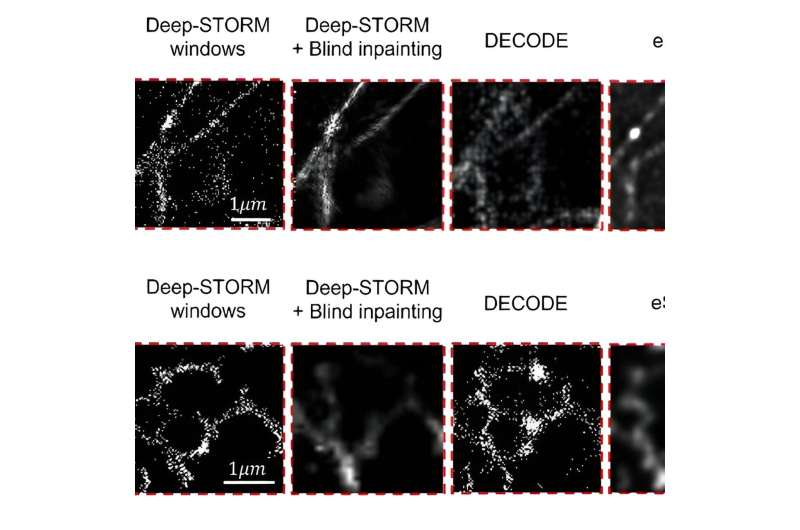
Using this methodology, the group was able to visualize multiple cell structures and their natural movement. They achieved a resolution of 30nm and temporal resolution of 15ms—an improvement of four orders of magnitude in the temporal resolution relative to the original SMLM method.
This new technology constitutes a major leap in biologists' ability to study living cells, a tool that will enable them to make new discoveries.
The study was done in collaboration with Dr. Onit Alalouf and Nadav Opatovski from the Technion, and Prof. Mike Heilemann and Soohyen Jang from Goethe University, Frankfurt.
More information: Alon Saguy et al, DBlink: dynamic localization microscopy in super spatiotemporal resolution via deep learning, Nature Methods (2023). DOI: 10.1038/s41592-023-01966-0
Journal information: Nature Methods
Provided by Technion - Israel Institute of Technology