This article has been reviewed according to Science X's editorial process and policies. Editors have highlighted the following attributes while ensuring the content's credibility:
fact-checked
peer-reviewed publication
trusted source
proofread
Scientists reveal machine learning could accelerate discovery of antimalarial properties in plants
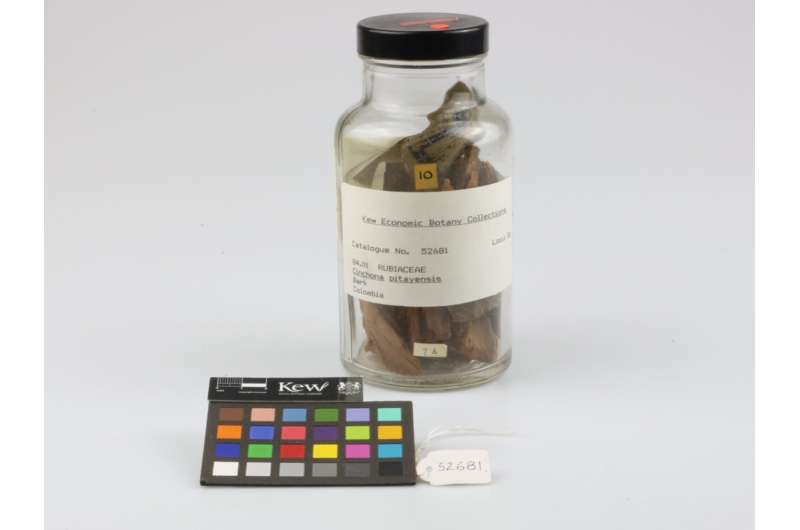

A new study, published on May 25, 2023 in Frontiers in Plant Science by researchers at the Royal Botanic Gardens, Kew and partners, has identified that the use of machine learning could speed up the search for plants with antimalarial properties.
Malaria is one of humankind's deadliest diseases and remains a significant global public health challenge. Malaria is caused by the Plasmodium parasite, which is spread to humans through the bite of infected mosquitoes. In 2021, the World Health Organization found that there were an estimated 247 million cases of malaria worldwide. Resistance to existing antimalarial drugs is an escalating challenge for eliminating malaria and, as a result, the WHO recommends that research into antimalarial medicines should be accelerated as part of an effort to reach global malaria targets.
Plants have provided or inspired the development of numerous pharmaceutical drugs as they are a rich source of bioactive compounds. For example, two major pharmaceutical drugs used in the treatment of malaria, quinine and artemisinin, are derived from plants. However, considering that there are an estimated 343,000 vascular plant species, identifying plants containing anti-Plasmodium compounds can be time-consuming and costly.
As part of the new study, scientists aimed to assess whether machine learning models could be trained on plant trait data to predict the anti-Plasmodium activity of plants. To do this they studied three flowering plant families—Apocynaceae, Loganiaceae and Rubiaceae—together constituting 21,100 species. Researchers explored an array of methods to demonstrate the effectiveness of machine learning algorithms and compared their performance with other approaches often used to select plants to find sources of bioactive compounds.
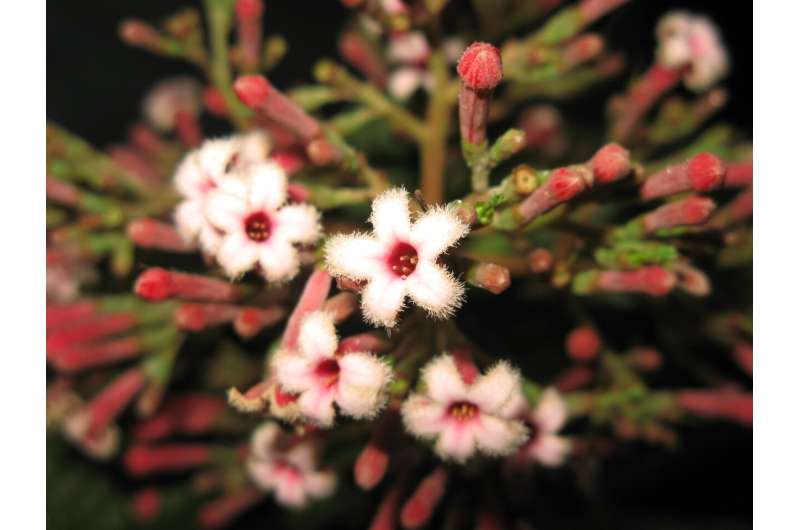

The experts found that the techniques provide a promising approach to improve the ability to predict plants with anti-Plasmodium properties, which could greatly accelerate the search for new compounds of pharmaceutical interest. Researchers estimate that 7,677 species in the three families warrant further investigation and that at least 1,300 active anti-Plasmodium species may have been missed using conventional approaches.
Adam Richard-Bollans, Research Fellow at RBG Kew, said, "Our results highlight the vast unexplored potential of plants to produce novel medicines. There are an estimated 343,000 known vascular plant species that remain largely unexplored scientifically, and we hope that our machine learning approach can be employed to efficiently search these plants for new medicinal compounds. These results also reinforce the importance of protecting biodiversity and using natural resources sustainably in order to preserve this valuable resource."
Daniele Silvestro, University of Fribourg and SIB Swiss Institute of Bioinformatics Group Leader, said, "Our study shows that machine learning offers the tools to combine scientific knowledge of plants and their traditional uses into a powerful predictive framework that can guide future testing and research. Biodiversity likely holds solutions to existing and future global health issues, and machine learning coupled with continued rigorous research in biology can help unlocking this potential."
More information: Adam Richard-Bollans et al, Machine learning enhances prediction of plants as potential sources of antimalarials, Frontiers in Plant Science (2023). DOI: 10.3389/fpls.2023.1173328
Journal information: Frontiers in Plant Science
Provided by Royal Botanic Gardens, Kew