New study indicates how deep learning can improve gene therapies and antiviral drugs
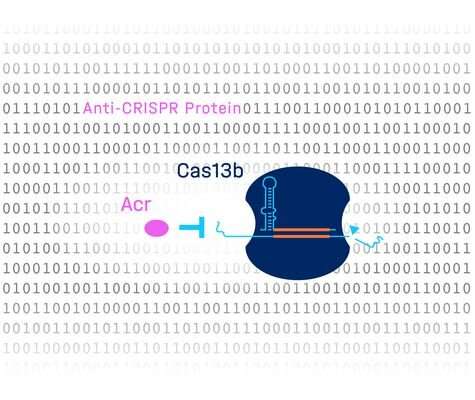
The nuclease Cas13b associated with CRISPR defense systems—also known as genetic scissors—has the potential to be used in the future in hereditary diseases to silence adverse genes. In the fight against infections, it is also being researched as an antiviral agent, as Cas13b can target the genome of viruses and render them harmless. Despite these promising features, researchers are looking for nuclease inhibitors that can control or stop such activities. The aim is to increase the safety and efficacy of future therapies and to help prevent off-target effects.
An international team led by the Helmholtz Institute in Würzburg, in cooperation with the University of Freiburg, has now applied deep learning for the first time to find natural nuclease inhibitors. The study, published today in the scientific journal Molecular Cell, identified the first such inhibitor that blocks the activity of Cas13b.
Bacteria often contain CRISPR-Cas defense systems that protect against viral invaders called phages. In response, phages have evolved proteins that can shut down CRISPR. These are called anti-CRISPR proteins, or Acrs. Researchers believe that there are numerous Acrs that have not yet been discovered. One of those researchers is Chase Beisel, head of the RNA Synthetic Biology Department at the Helmholtz Institute for RNA-based Infection Research (HIRI) in Würzburg, a site of the Helmholtz Centre for Infection Research (HZI) in Braunschweig in cooperation with the Julius-Maximilians-Universität (JMU) Würzburg. "Acrs can be used to better control CRISPR technologies," Beisel says, while acknowledging, "Identifying them means finding a needle in a haystack, since they look nothing like each other." This, he says, has led scientific studies in the past to come up with new prediction tools. However, the success of these methods has been limited, says Beisel. He initiated the study published today in Molecular Cell.
First example of using deep learning for Acr discovery
Consequently, a team of researchers from the HIRI in Würzburg, together with scientists from the Signalling Research Centres at the University of Freiburg and the Information and Computer Science Department at King Fahd University in Saudi Arabia, has pioneered the use of artificial intelligence to identify new Acrs.
"Using complex computational learning in combination with a high-throughput screen, we have succeeded in discovering a new anti-CRISPR protein that inhibits the clinically interesting nuclease Cas13b," says Katharina Wandera. The Ph.D. student in Chase Beisel's lab is working on the characterization of Acrs and is a first author of the current paper.
"The millions of predictions made by our algorithm open up completely new avenues for research," asserts Rolf Backofen from the University of Freiburg. His group developed the method, called DeepAcr, for the study.
The findings also suggest that DeepAcr can not only be used to discover new proteins that inhibit CRISPR-Cas systems lacking known Acrs. "The identified anti-CRISPR protein AcrVIB1 was not only unknown until now; it also seems to use a completely new mechanism of action," summarizes Omer Alkhnbashi, an assistant professor at King Fahd University who shares the first authorship with Wandera.
Characterizing this mechanism of action is one of the next steps that will now follow the scientific work.
More information: Katharina G. Wandera et al, Anti-CRISPR prediction using deep learning reveals an inhibitor of Cas13b nucleases, Molecular Cell (2022). DOI: 10.1016/j.molcel.2022.05.003
Journal information: Molecular Cell
Provided by Helmholtz Association of German Research Centres