New tool untangles complex dynamics on hypergraphs
Networks are a powerful model for describing connected systems in biological, physical, social, and other environments. As useful as they are, though, conventional networks are static and are limited to describing links between pairs of objects; they can't capture more complicated connections, like those that connect many points at once or those that change over time.
For systems built on more complex relationships, researchers turn to more sophisticated models like hypergraphs (which can show connections among groups), temporal networks (where connections change over time), or multilayer networks (which can show different kinds of connections). Connections among neurons in the brain, for example, change over time due to plasticity, and they include both electrical and chemical interactions.
But these generalizations come at a price, says physicist and SFI Schmidt Science Fellow Yuanzhao Zhang. "They're better at explaining realistic interactions in complex systems," he says, "but they're also more difficult to analyze because of that complexity."
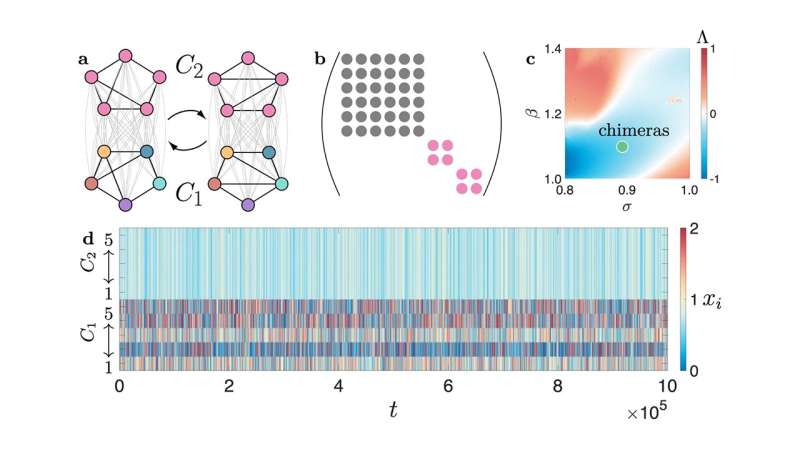
Difficult, but not impossible. In his work, Zhang focuses on finding ways to subdivide those thorny analytical problems, efficiently and optimally, into smaller, more manageable problems. Recently, he's been particularly interested in cluster synchronization patterns, which are a class of collective behavior that can emerge when subgroups in a network produce internally coherent but mutually independent dynamics. The cluster synchronization of certain frequencies of brain waves, for example, has been connected to improved memory.
In a paper published recently in Communications Physics, he and his collaborators—including SFI science board member and Northwestern University physicist Adilson Motter—describe a new framework for simplifying the analysis of synchronization patterns in a wide variety of systems that include hypergraphs, temporal networks, and multilayer networks. Their approach uses a technique for simultaneously simplifying multiple matrices, which can be applied to large (generalized) networks with tens of thousands of nodes.
Zhang says the technique could help researchers investigate new phenomena that only emerge in the presence of higher-order interactions, described by models like hypergraphs. It might also be useful in understanding systems with both pairwise and non-pairwise interactions. "There could be some synergy between those interactions that we don't fully understand," he says. "When you have a system like that, how do those two types of interactions interact with each other? It's an interesting question."
More information: Yuanzhao Zhang et al, Unified treatment of synchronization patterns in generalized networks with higher-order, multilayer, and temporal interactions, Communications Physics (2021). DOI: 10.1038/s42005-021-00695-0
Journal information: Communications Physics
Provided by Santa Fe Institute