CRISPR tech could enable early diagnosis of devastating citrus disease

Penn State and U.S. Department of Agriculture scientists have used cutting-edge CRISPR/Cas technology to develop a diagnostic test that could enable early diagnosis of citrus greening, or Huanglongbing, a serious disease that threatens worldwide citrus production, which is valued at roughly $17 billion from the sale of fresh fruit and juices.
In a study newly published in the journal Phytopathology, the researchers demonstrated that the new assay can detect the presence of the disease's causal agent—the bacterium Candidatus Liberibacter asiaticus, abbreviated as CLas—at a sensitivity level 100 to 1,000 times greater than a commonly used diagnostic test, quantitative polymerase chain reaction, or qPCR.
Citrus greening, sometimes referred to as HLB, was described in Asia more than a century ago and reached Florida in 2005. Since then, the disease has decimated that state's orange crop, reducing production by more than 70%. The pathogen also has spread to Texas, California, Georgia and Louisiana, threatening the $3.35 billion U.S. citrus industry.
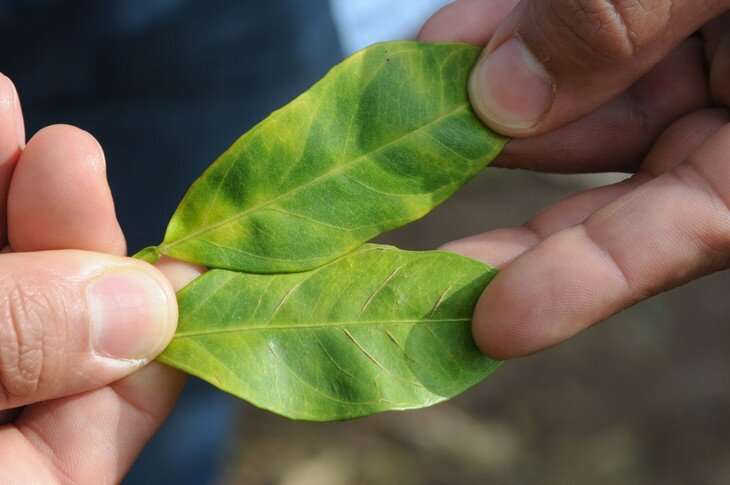
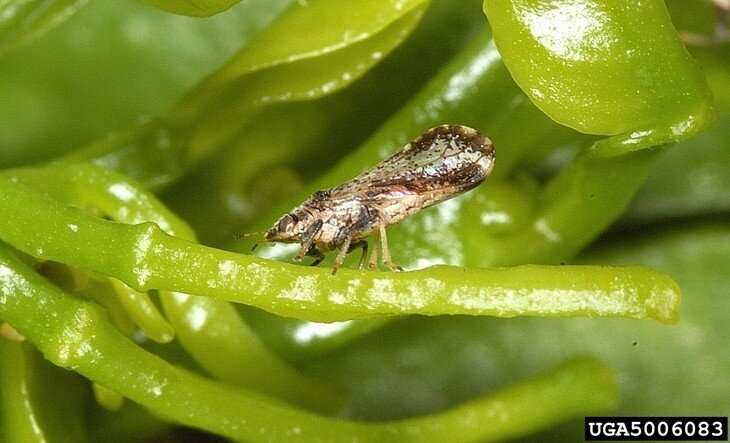
Spread by an insect known as the Asian citrus psyllid or the grafting of infected tissues, CLas does not harm people or animals, but once a citrus tree is infected, there is no cure for the disease. According to USDA, infected trees display asymmetrical yellowing of the leaves and leaf veins—referred to as "blotchy mottle"—and produce fruit that is green, misshapen and bitter, unsuitable for the fresh fruit or juice markets. Most infected trees die within a few years.
Scientists say the best hope of reducing the spread of citrus greening is to eliminate diseased trees quickly. As a result, early detection of the pathogen is crucial because infected trees can act as a disease reservoir for months or years before showing visible symptoms.
To address the need for early diagnosis, researchers deployed CRISPR/Cas, a powerful gene-editing technology that recently has been adapted as a molecular diagnostic tool, noted study co-author Yinong Yang, professor of plant pathology in Penn State's College of Agricultural Sciences.
"So far, various technologies, such as canine olfactory detection and qPCR, have been used to diagnose and confirm CLas infection," said Yang, who also is affiliated with the Huck Institutes of the Life Sciences. "But these tools are often inadequate for early detection of CLas in asymptomatic tissue—where low amounts of bacteria are present—and are unsuitable for high-throughput diagnosis in the field."

CRISPR stands for clustered regularly interspaced short palindromic repeats. The technology can modify an organism's genome by precisely delivering a DNA-cutting enzyme—Cas—to a targeted region of DNA. The resulting modification can delete or replace specific DNA pieces, thereby promoting or disabling certain traits. CRISPR/Cas has many potential biomedical and agricultural applications, such as treating genetic diseases or developing hardier crops.
The researchers adapted the CRISPR/Cas system to develop an assay based on a platform known as DETECTR, which stands for DNA endonuclease-targeted CRISPR trans reporter. In this assay, a Cas variant called Cas12a is mixed with CRISPR RNA designed to seek out CLas DNA, and a synthetic molecule known as a fluorescent reporter oligonucleotide. If CLas DNA is present, the Cas12a cleaves both the pathogen's DNA and the reporter oligonucleotide, enabling the generation of a fluorescent signal for detection.
The researchers report that this assay was highly sensitive and specific in detecting CLas DNA, using either a microplate reader for fluorescence readouts or a lateral flow strip for a visual result indicating the presence of a target sequence—similar in concept to a home pregnancy test.
"The assay successfully detected and confirmed the presence of CLas nucleic acids in infected samples collected from Florida," said study lead author Matthew Wheatley, postdoctoral scholar in plant pathology and environmental microbiology, Penn State.
"We tested DNA samples from infected sweet orange, pomelo, grapefruit and periwinkle and from the citrus psyllid vector, as well as from uninfected grapefruit and psyllid samples as negative controls," Wheatley said. "The CLas DETECTR assay confirmed that all infected samples were positive for CLas compared to the negative controls. Interestingly, two grapefruit DNA samples that previously tested CLas negative using a qPCR test showed weak positivity for CLas in the DETECTR assay."
Yang pointed out that the DETECTR assay's compatibility with lateral flow technology holds promise for providing rapid and economical testing for HLB in the field.
"These results show that CRISPR/Cas-based detection strategies have the capability to improve upon current diagnostics of citrus greening by providing specific and highly sensitive detection of CLas nucleic acids," he said. "The improvement in detection sensitivity and ease of use compared to traditional methodologies like qPCR position the CLas DETECTR assay as a promising tool for HLB diagnostics in regions where the CLas pathogen occurs."
More information: Matthew Wheatley et al. Highly sensitive and rapid detection of citrus Huanglongbing pathogen (Candidatus Liberibacter asiaticus) using Cas12a-based methods, Phytopathology (2021). DOI: 10.1094/PHYTO-09-20-0443-R
Provided by Pennsylvania State University