Discovery of pH-dependent 'switch' in interaction between pair of protein molecules
All biological processes are in some way pH-dependent. Human bodies and those of other organisms need to maintain specific and constant pH regulation in order to function. Changes in pH can have serious biological consequences—or serious benefits, as researchers at the Qingdao Institute of Bioenergy and Bioprocess Technology (QIBEBT), Chinese Academy of Sciences (CAS) found.
The findings are published on Oct. 23 in the journal Science Advances.
Cellulosomes are extracellular complexes consisting of multiple enzymes, which are associated with the cell's surface. Within the cellulosome cellular structure, the protein molecules dockerin and cohesin were the focus of this study.
"Cellulosomes are complex nanomachines in nature and have great values in biofuel production and biotechnology. This study is an example of the complexity and diversity of cellulosomes," said study author Feng Yingang, Professor, Metabolomics Group.
Changes in pH have previously been shown to result in "on-off" switches within protein functions, many of which occur naturally and are essential for life processes. Biotechnical innovations can utilize this relevant phenomenon to develop sensors or switches using biomolecules that are pH-dependent.
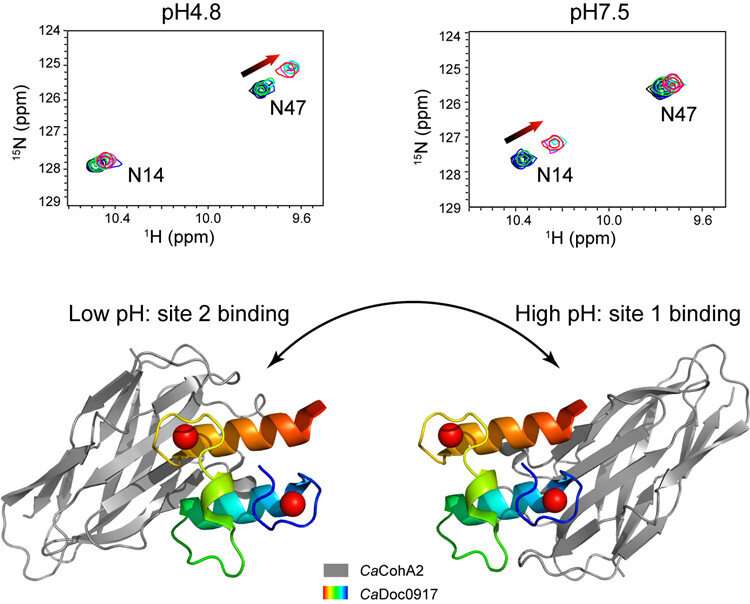
The latest discovery, on the cellulosome assembly of the bacterium Clostridium acetobutylicum, takes this prospect further by switching between two functional sites, rather than simply on or off. This opens additional possibilities.
"Our study not only revealed an elegant example of biological regulation but also provides a new approach for developing pH-dependent protein devices and biomaterials for biotechnological application," said Feng.
Researchers found that changing the pH from 4.8 to 7.5 resulted in the cohesin-binding sites on the dockerin molecule switching from one site to the other. This type of switching between two functional sites has not been noted for any interaction between proteins previously.
Nuclear magnetic resonance (NMR) and isothermal titration calorimetry (ITC) were used to describe the distinct features of this interaction. Researchers additionally noted that the affinity, or the attraction between the molecules, was found to change along with the pH. This property is considered unusual when compared to other cohesin-dockerin interactions and is unique, thus far, to C. acetobutylicum bacteria.
These, and future discoveries like it, can potentially be used to create more complex biological switches in synthetic biology and further developments in the fields of biotechnology.
"Next, we will continue to elucidate the structure and regulation of cellulosomes, which could provide interesting novel discoveries and new strategies to increase the efficiency of lignocellulose-based biofuel production," Feng said. "Our ultimate goal is to promote sustainable and economical lignocellulose bioconversion and bioenergy production."
More information: "Discovery and mechanism of a pH-dependent dual-binding-site switch in the interaction of a pair of protein modules" Science Advances (2020). DOI: 10.1126/sciadv.abd7182
Journal information: Science Advances
Provided by Chinese Academy of Sciences




















