Studying gene function in animal models
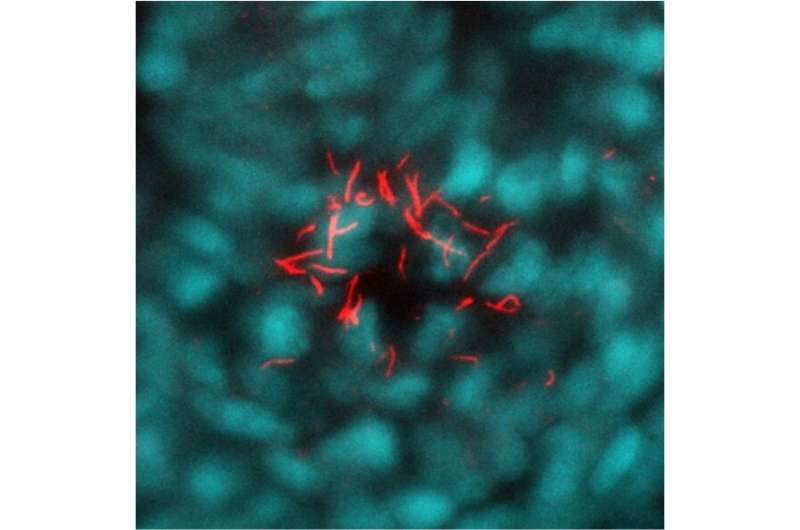
Researchers from the group of Jeroen Bakkers have described a thorough way to study the function of genes in model animals such as the zebrafish. In their study, published in Nature on September 23rd, they give an example of such a comprehensive study into gene function and provide a guideline to fellow researchers.
All organisms, from bacteria to plants and animals, have a very large number of genes in their DNA that together determine everything about that organism: from how it develops to the color of its eyes or leaves. Biologists have long been interested in finding out the functions of all these individual genes to gain understanding of how organisms work, how they develop, how genetic diseases originate and how they may be treated. One of the organisms used in such fundamental biological research is the zebrafish, a small tropical fish that is used as a model for vertebrate development.
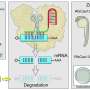
Genes in the DNA are like master instructions for making proteins. Because the master instructions need to remain safe, small copies of the genes are made in a molecule called RNA—you could compare it with printout of the large database of instructions that is the DNA. Such a copy is then used in the cell as an instruction to make proteins; like you would use an IKEA instruction manual to build a cabinet.
Knock-down and knock-out
There are multiple ways to study the function of a gene: the gene may be disabled in different ways in an organism such as the zebrafish, which is then compared to a normal, or "wild-type" zebrafish. Referring back to the database analogy, one of the ways to disable the function of a gene is to block all the printouts of the instructions, or RNA molecules, so that no more proteins are built—like intercepting all copies of instruction manuals for a certain cabinet to make sure nobody can build that cabinet anymore.
This method is called "knock-down" and uses a morpholino, a small molecule that blocks the RNA molecule, the printout, from being used to build proteins. Another way to disable the function of a gene is to "knock-out" the gene in the DNA itself, by creating change, or a mutation in the DNA. In our analogy this would be to change, or even delete, the folder containing the original instructions in the master database of IKEA, so that no more printouts, or RNA molecules, can be made, and no cabinets, or proteins can be built.
Knock-down approaches such as morpholinos have been widely used in many organisms, including zebrafish, to study gene function. These approaches are quick, and the effects of morpholinos specifically are transient—once the morpholino is gone, printouts can be used freely again. Changing the DNA itself—as done in knock-out models—proves more difficult, and permanent, but ensures that no more printouts are made.
Discrepancies
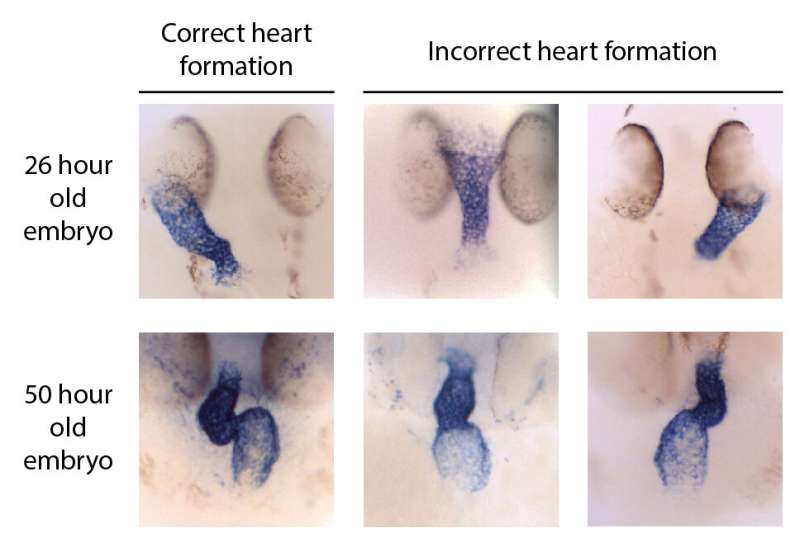
Sometimes, the results from knock-down studies and knock-out studies do not match, which raises the question which results are correctly describing the function of the gene that was knocked-down or knocked-out. In collaboration with the groups of Gage Crump (University of South California) and Jamie Nichols (University of Colorado), researchers from the group of Jeroen Bakkers describe a thorough way to address these discrepancies, as they came across a study that described how a gene, prrx1a, was involved in making sure that the formation of the heart occurred properly.
In this study, a morpholino was used to inhibit prrx1a, which resulted in mistakes in the formation of the heart, leading to the conclusion that prrx1a plays a role in heart development. However, zebrafish in which this gene was disabled in the DNA—a knock-out for the gene prrx1a—were available and did not show mistakes in heart formation. This prompted the researchers of Bakkers' group to investigate the function of the prrx1a gene in more detail.
Compensation
An additional level of complexity in the study of knock-outs was unraveled recently, with the description of what is known as transcription adaptation. When knocking out a gene, we often actually introduce mistakes in its sequence, so that the produced protein is not functioning properly. The organism, however, detects this and manages to regulate the activity of related genes to compensate. Using the cabinet analogy: if a printout of an instruction booklet is incorrect or incomplete, the booklet of another cabinet can be used to complete building it.
This mechanism may be circumvented if printout instructions are altogether missing, for example by intercepting RNA through the use of a morpholino or by preventing the mere existence of the targeted RNA. The latter technique was applied by the researchers for the current study. They made various so-called RNA-less prrx1a zebrafish—zebrafish in which no RNA molecules or printouts are produced—by deleting the entire prrx1a locus or its promoter region and part of its coding sequence. None of these techniques resulted in mistakes in heart formation.
Need for validation
Furthermore, the researchers used a morpholino for the prrx1a gene in zebrafish in which the prrx1a gene was already knocked out in the DNA. These zebrafish do not make any RNA, or printouts, of the prrx1a gene, so a morpholino that intercepts these RNA molecules should not have any effect in these fish. However, when the researchers used the morpholino in the prrx1a knock-out zebrafish they saw the same mistakes in heart formation. This shows that the mistakes in heart formation ensue from the presence of the morpholino itself, and not from specific inhibition of the function of the prrx1a gene.
With this study, the researchers show that investigating the function of a gene should be done carefully, especially in the case of inhibitors such as morpholinos that intercept the RNA molecules. The results should always be validated in multiple ways to show that the method itself is not responsible for the results that the researchers find.
More information: Federico Tessadori et al. Zebrafish prrx1a mutants have normal hearts, Nature (2020). DOI: 10.1038/s41586-020-2674-1
Journal information: Nature
Provided by Hubrecht Institute