A race to solve the COVID protein puzzle
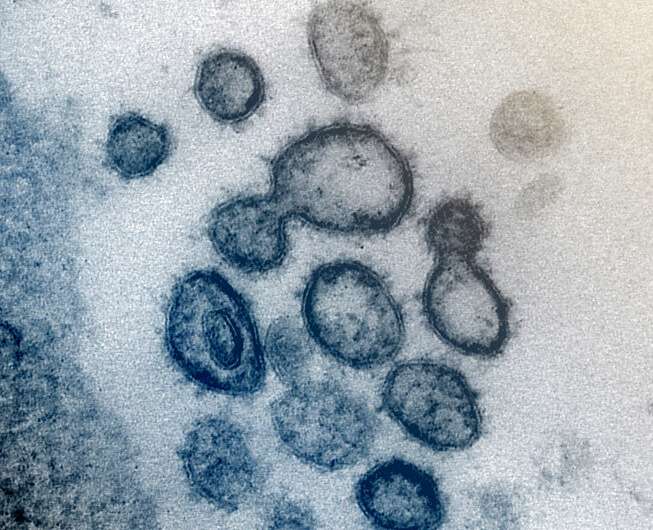
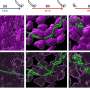
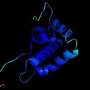
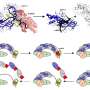
Among the many unknown factors in the science of COVID-19, one involves the structures of the proteins that make up the exterior of the coronavirus. A coronavirus particle has multiple proteins, including the familiar spiky structures on the outside of the spherical virus particle (aptly named "spike proteins"). While researchers know the specific amino acids that make up the various proteins, their three-dimensional structures are not known.
Corey O'Hern, professor of mechanical engineering & materials science, physics, & applied physics, is among the researchers trying to determine their structures. To do so, he uses machine-learning techniques that can determine whether a computational model of a protein is accurate or not (even without knowing the actual experimentally determined structure of the protein).
Along with other researchers around the world, O'Hern will submit his findings to the Critical Assessment of Protein Structure Prediction (CASP), a competition among researchers in the field to predict the structure of proteins that have not yet been characterized experimentally. A few months ago, CASP's organizers put out a call for studies of the coronavirus proteins to better understand their structures.
"They posted computationally generated structures for these COVID proteins, so now there's a race to determine whether any of these potential structures are correct," O'Hern said.
What's the benefit of determining these COVID protein structures?
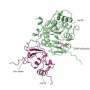
The structures of these proteins are related to the ability of the coronavirus to bind to cells and invade them, so knowing their structures means that you may be able to design a drug to bind to the protein and prevent the virus from invading the cell. One of the fundamentals for drug design is knowing the structure of the protein so that you can design a complimentary ligand that will bind to it and inhibit the original function of the protein.
How do you predict the structures of these proteins?
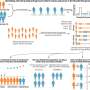
That's the hard thing. The main way that people experimentally solve protein structures is through crystallization. It is kind of black magic—you never know which proteins will crystallize and under what conditions. COVID proteins are difficult to crystallize. So, the goal in the O'Hern group is to first understand the structure of the more than 100,000 experimentally solved protein structures, and then determine whether the computational models of COVID proteins have the same features as those of the current library of known protein structures. If the models do have the correct features, we can use molecular dynamics simulations to guide the computational models toward more experimentally reasonable structures.
How do you know when you get it right?
Unfortunately, you don't know for sure whether you have the correct structure of the COVID proteins until you can crystallize them, carry out nuclear magnetic resonance experiments, or otherwise experimentally characterize them. However, many scientists around the world are submitting computational models to CASP. We can evaluate all of them using our machine learning approach to determine whether they are physically reasonable. We can also take the best computational models and begin work on drug design.
Provided by Yale University