New tool could help scientists dig deeper for prized crop genes
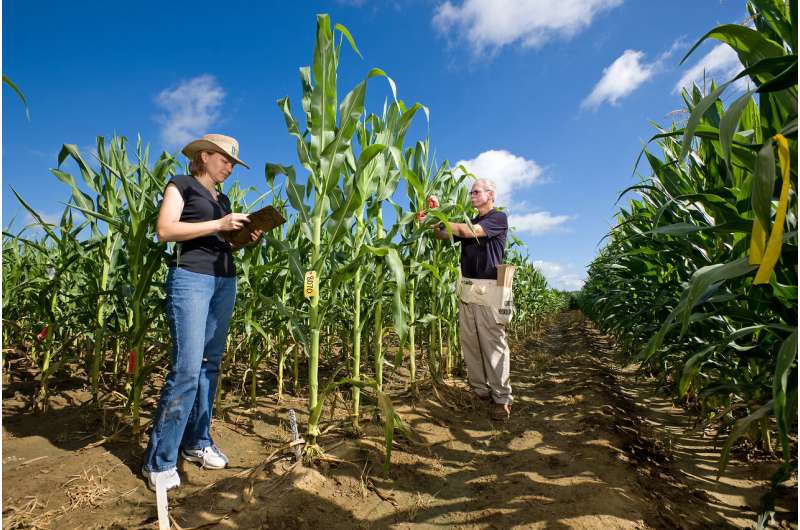

A new computer application (app) from Agricultural Research Service (ARS) scientists could speed the search for genes that underpin important crop traits, like high yield, seed quality and resistance to pests, disease or adverse environmental conditions.
Known as the Pathway Association Studies Tool (PAST), the app allows users to build on the results of genome-wide association studies (GWAS) of crops. GWAS takes a kind of bird's-eye look at a crop plant's genome for marker regions called single nucleotide polymorphisms (SNPs). Finding SNP markers near the gene or genes encoding a desired trait can flag the genomic whereabouts of those genes, and also help plant breeders follow the trait's inheritance and expression. This makes it easier to select plants that have the desired trait and develop new, elite varieties from them for producers.
However, GWAS's use of a statistical threshold means that only markers with the strongest gene associations are identified, noted Marilyn Warburton, a geneticist with ARS's Corn Host Plant Resistance Research Unit (CHPRRU) in Mississippi State, Mississippi. This can blind researchers to the presence of other gene markers that fall below the threshold, but are no less meaningful to their studies, she added.
PAST picks up the slack by taking an additional step known as metabolic pathway analysis. This added step doesn't just find interesting markers that GWAS misses; it also reveals important biological information about their associated genes and how each contributes to the biochemical assembly of a plant trait, function or response—regardless of the crop species.

Warburton's team published a paper describing PAST in the January 2020 issue of the journal Plants and posted an announcement about the app to her LinkedIn account, which has received 5,500 views to date. About 900 users from around the world have downloaded the app so far, and Warburton hopes for even greater awareness of it following the March 12 release on the MaizeGDB website, a database with extensive genomics and genetics information about corn (maize).
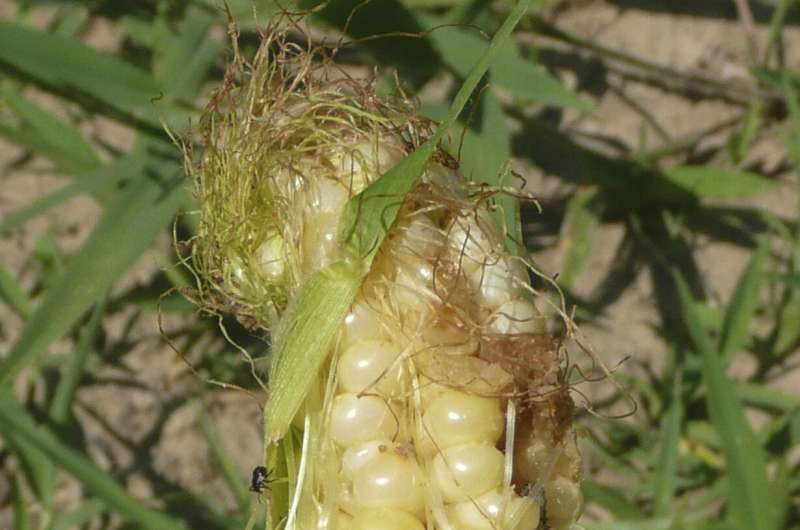
In her own research, Warburton's use of both GWAS and PAST has already led to the identification of genes in corn plants for resistance to the corn earworm (a caterpillar pest) and Aspergillus flavus, a greenish mold that produces a carcinogen called aflatoxin. Unchecked, corn earworms feed on the corn plant's silks and kernels, causing damage that fosters Aspergillus growth and aflatoxin contamination.
By federal law, corn or other grains with aflatoxin levels exceeding 20 parts per billion cannot be sold for human consumption, and the grains' use for animal feed is restricted. In the United States, outbreaks of Aspergillus molds that produce the carcinogen inflict more than $200 million annually in economic losses for corn and $300 million for peanuts and other crops combined. Warburton's research is part of a broader effort at CHPRRU together with Mississippi State University (MSU) collaborators to pre-empt aflatoxin on multiple fronts—with plantings of resistant corn varieties being a keystone defense.
She credited Adam Thrash (MSU graduate student), Daniel Peterson (MSU faculty), Mason Deornellis (MSU undergraduate) and Juliet Tang (an ARS postdoc, now a scientist with the Forest Service) with assisting in the development, testing and release of PAST.
Warburton expects the app's ease of use and versatility could also buttress GWAS studies of animals and even humans, in which the identification of genes linked to inherited diseases can lead to personalized medicines to treat these diseases.
More information: Adam Thrash et al. PAST: The Pathway Association Studies Tool to Infer Biological Meaning from GWAS Datasets, Plants (2020). DOI: 10.3390/plants9010058
Provided by Agricultural Research Service