Metabolic fossils from the origin of life
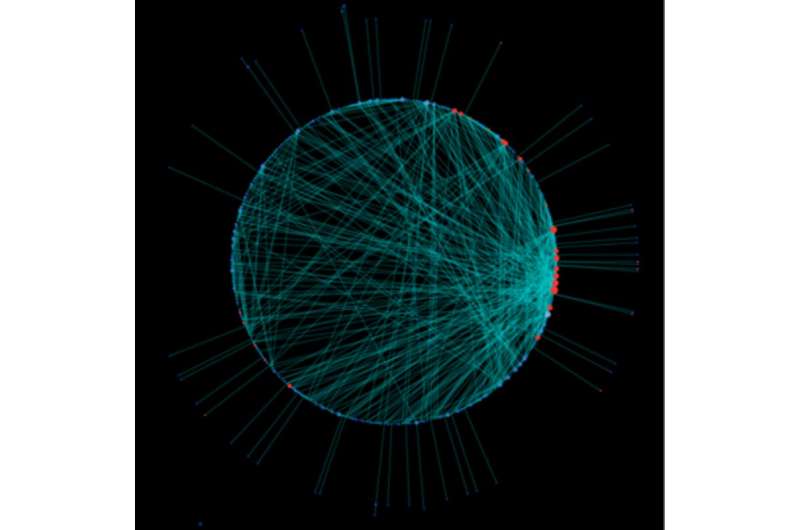

Life converts food into cells via dense networks involving thousands of reactions. New research uncovers insights as to how such networks could have arisen from scratch at life's origin. An international team of researchers in Germany, New Zealand and the U.S.A. has investigated metabolic networks of primitive microbes and identified autocatalytic sets (interconnected collections of self-reinforcing reactions) that are older than genes.
Living cells are the end product of metabolic networks. Food molecules that enter the cell are converted to central intermediates that are then channeled into the pathways that produce the molecules of which cells are made. These networks typically entail more than 1000 reactions, almost all of which are performed by enzymes (proteins), which are encoded by genes (nucleic acids). The link between genes and proteins is, in turn, the universal genetic code that instructs ribosomes to make proteins according to the information stored in genes.
These components are all interlinked: the ribosome is 50% protein and 50% RNA by weight, the proteins are made of amino acids, the RNA is made of nucleic acid bases, and the amino acids and bases are made by the roughly 1000 reactions in metabolism, which are catalyzed by the enzymes that the genes encode. With so many layers of mutual interdependence, it is no wonder that scientists have been flatly stumped for over a century when it comes to the question of how such a complex system of interactions could arise at the origin of life. As with the evolution of all complex systems, it had to start from something simpler. But what? New findings by Joana C. Xavier and colleagues reported in Proceedings of the Royal Society B in London provide new inroads into this longstanding question.
The new clues come from the least expected of all places: mathematics. Almost 50 years ago the American polymath Stuart Kauffman suggested that theoretical constructs called autocatalytic sets might have been intermediates in the origin of molecular complexity of the kind that we find in metabolism and cells. Such autocatalytic sets consist of elements (members of the set) that are both products and catalysts such that they can make more of themselves given suitable starting material. The analogy to metabolism and enzymes is evident.
The existence and properties of such autocatalytic sets remained the subject of much speculation and decades of fierce debate until the mathematician Mike Steel from the University of Canterbury in New Zealand and Wim Hordijk, a computer scientist from The Netherlands, both coauthors on the study, found ways of harnessing them in the computer. They found that a particular class of autocatalytic sets called RAFs (for reflexively autocatalytic food generated networks), which are very similar in design to cellular metabolism, have the unexpected property of being downright likely to arise from scratch. "The surprise is that the elements only need to add a tiny amount of catalysis to the system before they start to make more of themselves," says Steel. "This is what physicists call self organization, a kind of holy grail in origin of life research," adds Hordijk.
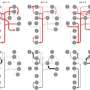
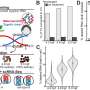
With a background in the metabolic networks of real cells, Joana C. Xavier in the Institute for Molecular Evolution at the University of Düsseldorf asked whether RAFs could be detected in the metabolic networks of the most primitive microbes, strict anaerobes that live from H2 and CO2. Indeed, she found that RAFs were there in the metabolism of ancient anaerobes, but they were substantially smaller than the whole metabolic map, comprising only 394 reactions in the case of an ancient microbe that converts H2 and CO2 to acetate for a living, and 209 reactions in the case of an ancient microbe that converts H2 and CO2 to methane. "This intermediate size is interesting," says Xavier, "because it points to an intermediate state in the evolution of metabolism, something more complex than individual reactions but less complex than a cell."
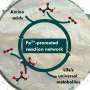
The two kinds of unicellular organisms at the focus of the study, called acetogens and methanogens, have long been in the sights of microbiologists interested in the origin of life. They have been linked to the last universal common ancestor, LUCA, and to geochemical reactions at hydrothermal vents. Xavier found that the acetogen and methanogen sets overlap to form an ancient core network of 172 reactions. This ancient conserved core predates the divergence of bacteria and archaea and has intriguing properties. It can generate amino acids and nucleic acid bases from a simple starting food set, but if provided only the bases as food, no network at all emerges. "Not only have autocatalytic networks left fossils in real metabolism, they preceded both RNA and protein polymers in evolution, that is a step forward in my book," says Kauffman, coauthor of the study and autocatalysis pioneer.
William Martin at the University of Düsseldorf and coauthor of the study says "The networks that trace to LUCA's metabolism are older than genes, they point to natural order in the chemical reactions of life." Acetogens and methanogens grow under the kinds of conditions that are encountered today at hydrothermal vents. Did life arise at hydrothermal vents? "The closer we look, the more signs keep pointing in that direction" says Xavier, "the idea keeps uncovering findings that converge. These vents were probably the first bioreactors on Earth."
The identification of autocatalytic networks as components of modern metabolism takes them off the drawing board and into the real world of microbial life. That they reveal fossils from the earliest stages of chemical evolution was unexpected, and opens up new routes for the study of our deepest evolutionary past, probing the time 4 billion years ago, when life was just starting from a small set of naturally-occurring chemical reactions that took place somewhere, perhaps at a hydrothermal vent.
More information: Joana C. Xavier et al, Autocatalytic chemical networks at the origin of metabolism, Proceedings of the Royal Society B: Biological Sciences (2020). DOI: 10.1098/rspb.2019.2377
Journal information: Proceedings of the Royal Society B
Provided by Heinrich-Heine University Duesseldorf