New diagnostic tool developed for global menace Xylella fastidiosa increases specificity

The bacterium Xylella fastidiosa is notable for having a wide host range, with the ability to infect more than 300 plants. X. fastidiosa has a long history of causing serious harm to crops and trees in the Americas, with especially damaging repercussions on grapevine and citrus.
In 2013, X. fastidiosa was discovered for the first time outside of the Americas, attacking olive trees in southern Italy causing olive quick decline syndrome. Since then, it has been increasingly found in various environments throughout France and Spain on a variety of plant species and the demand for fast and reliable diagnostic tools is crucial to effective disease management strategies.
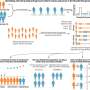
In a research article in Plant Disease, Bonants et al. record their efforts to improve the reliability of existing X. fastidiosa diagnostic tools. The team combined two existing tools with an internal control to develop a triplex TaqMan assay, which they then used to analyze DNA extracts in naturally infected plant material, artificially infected plant material, and uninfected plant material.
The triplex TaqMan assay has increased specificity as it targets two loci rather than just on locus on an X. fastidiosa genome. This is the first time a diagnostic tool of this type has been successfully developed for this pathogen.
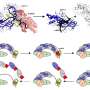
Additionally, the researchers developed procedures for analyzing DNA extracts from both infected and healthy plants using next-generation sequencing (NGS) technology, marking the first time extracts from X. fastidiosa-infected plants were analyzed in this way. In all samples, DNA reads were detected specific for X. fastidiosa and in most cases the pathogen could be identified to the subspecies level. This new procedure leads the way for future track-and-trace studies.
More information: Peter Bonants et al, Development and Evaluation of a Triplex TaqMan Assay and Next-Generation Sequence Analysis for Improved Detection of Xylella in Plant Material, Plant Disease (2018). DOI: 10.1094/PDIS-08-18-1433-RE
Provided by American Phytopathological Society