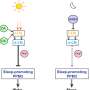
Why the system needs sleep
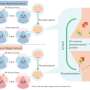
Sleep is essential for brain functionality and overall health but understanding how sleep delivers its beneficial effects remains largely unknown. In an article publishing on August 9 in the open access journal PLOS Biology, Shanaz Diessler and Maxime Jan at the University of Lausanne and the SIB Swiss Institute of Bioinformatics (Switzerland), sleep researchers are exploring new and unbiased approaches that can take sleep to a systems level. In one such approach, referred to as 'systems genetics', inferences about biological phenomena can be made by linking together several levels of information from DNA to phenotype via gene expression, proteins and metabolism at the level of a population. Systems genetics offers a global and interconnected view of biological phenomena and is therefore considered critical towards predicting disease susceptibility.
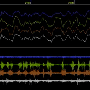
The study performed systems genetics analysis in mice with the aim of identifying the molecular signaling pathways that predict the resilience or susceptibility of sleep loss. Seven years of work culminated in an open access and interactive knowledgebase (see https://bxd.vital-it.ch/) quantifying the effects of sleep deprivation and of the genome at the level of the brain and liver transcriptome, of the blood metabolome, and, finally, of the sleep-wake phenome referring to an unprecedented detailed dissection of brain activity and sleep-wake behavior.
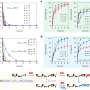
Initial exploration yielded several remarkable new insights. Preventing sleep during the first half of the habitual rest phase (the light period in the mouse), pervasively affected all systems levels with 78% of all expressed genes in the brain changing their expression. The authors identified a number of genetic variants that exerted their action only after the sleep deprivation challenge and not when left undisturbed. Moreover, these variants not only determined the magnitude of the molecular or sleep response to sleep loss but also, in some cases, the direction of the change, explaining the strong inter-individual changes in coping with insufficient sleep.
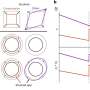
Perhaps the most striking observation that came from these analyses is that molecules and metabolic pathways active in peripheral tissue only (the liver) seem to directly influence phenotypes considered strictly central such as the frequency of oscillations known to originate from specific brain areas such as the hippocampus. Converging evidence drawing on all four levels of organization implicated fatty acid metabolism as a substrate of the negative consequences of sleep loss. Such findings not only can direct novel therapeutic strategies but also challenge the largely brain-centric view of sleep that has long guided hypotheses on its function and regulation.
The two groups led by Paul Franken and Ioannis Xenarios continue their collaboration and further explore and strengthen this unique resource by adding a third regulatory molecular layer and by developing the bioinformatic tools needed to bring to light the many interactions and pathways still hidden in the data. The authors hope that through this unbiased approach, we will learn why the system needs sleep.
More information: Diessler S, Jan M, Emmenegger Y, Guex N, Middleton B, Skene DJ, et al. (2018) A systems genetics resource and analysis of sleep regulation in the mouse. PLoS Biol 16(8): e2005750. doi.org/10.1371/journal.pbio.2005750
Journal information: PLoS Biology
Provided by Public Library of Science