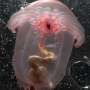
Scientists are using DNA to study ocean life and reveal the hidden diversity of zooplankton
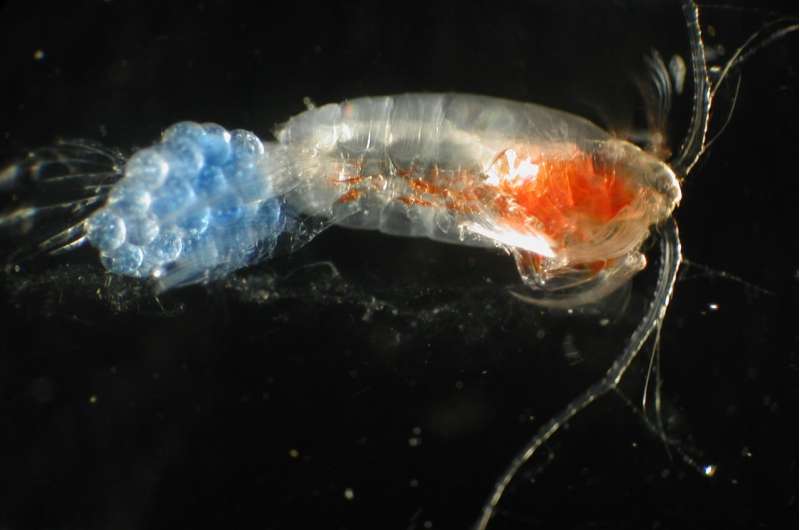

Marine zooplankton are tiny animals, roughly the size of insects you might see on a summer day, that drift with ocean currents. Many of them are lovely, but except for scientists who study them, few people are aware that they are among the most numerous – and important – animals on Earth.
My research
focuses on marine zooplankton, which I think of as "charismatic microfauna." These minute organisms are key players in open
More than 7,000 species of zooplankton have been described, but we do not know how many total species exist. Since they are important food sources for larger fish, and respond rapidly to environmental shifts and climate change, knowing more about zooplankton is key for understanding the health of ocean ecosystems.
My laboratory has gained new insights into zooplankton diversity using inexpensive and easy-to-use DNA sequencing. These approaches are providing new tools to identify species. According to some estimates, there could be as many as 70,000 unknown and overlooked zooplankton species yet to be discovered.
Key links in ocean ecosystems
The open ocean is the largest habitat by volume on Earth. Because marine zooplankton are so numerous and diverse, they help to create complex food webs with multiple pathways of who-eats-whom. But changes in environmental conditions, including global warming, can disrupt these systems by altering zooplankton diversity and distribution.

Currently, key zooplankton species are shifting northward in search of cooler waters in a warming ocean. This is causing major disturbances to ocean ecosystems. For example, the cod fishery in the North Sea crashed after the zooplankton species that was the fish's preferred prey was replaced by another closely related copepod species that moved north in response to global warming.
Events like this show that zooplankton can play critical roles in ocean ecosystems, even though scientists know very little about them, and may not appreciate the importance of a given species until it is gone.
Many factors make it challenging to measure marine zooplankton diversity. First, collecting samples from the deep sea is hard work. Research vessels operate around the clock, with scientists on board usually working two six-hour watches each day. We sample zooplankton using electronically operated, highly complex net systems.
When the net is recovered, the samples contain many different species and the animals are usually still alive. They are carefully moved into trays for observation and species identification. In spite of the demanding schedule and hard work, I have loved going to sea and have been on many cruises, including stunning explorations of the polar regions of the Arctic and Antarctic Oceans.
For my research, samples collected at sea are quickly preserved for genetic analysis, which is usually done back in the lab. This is painstaking work. Zooplankton as a group are taxonomically complex, meaning that they include many difficult-to-identify and rare species. Also, many important marine organisms, including fishes, lack diagnostic characteristics in their immature (larval) stages.
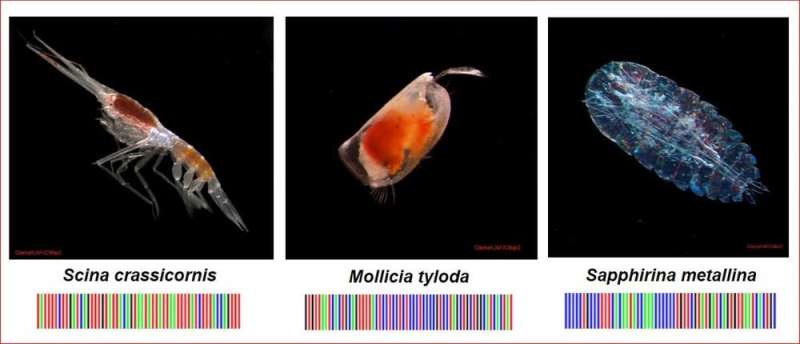
Identifying zooplankton species with genetic markers
To understand, assess and manage marine ecosystems, it is critical to have information about the diversity of species that make them up. But acquiring this information using traditional methods is tedious, expensive and slow.
The job falls to expert morphological taxonomists, who may spend years learning to recognize subtle characteristics that mark differences between species. Since there are 7,000 described species of zooplankton spanning 15 phyla and more than 30 different groups, this work can involve large teams.
But molecular methods are providing important new insights. My laboratory group uses different molecular approaches to study population ecology, population genetics, species diversity and other questions about marine zooplankton.
One efficient approach is to use DNA barcodes – short sequences of DNA – to identify and distinguish species. To do this, we sequence the DNA from a genetic region that is known to occur in all zooplankton, but varies markedly from species to species.
My laboratory has determined DNA barcodes for many zooplankton species. We use a portion of the mitochondrial cytochrome oxidase I (COI) gene, which is the most commonly sequenced gene region for analysis of species diversity among marine animals. The growing barcode database has been likened to a Rosetta Stone for species identification of global zooplankton. COI sequences for thousands of species of marine animals are now available. They create an invaluable reference library that can be used as a basis for next-gen environmental sequencing, also known as metabarcoding.
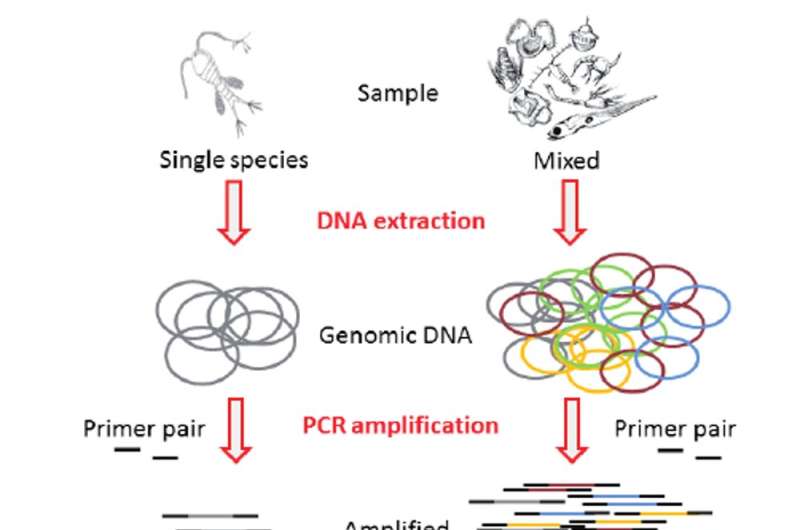
Using metabarcoding speeds up our work because we can sequence DNA from zooplankton samples without identifying individual specimens. This "high-throughput" DNA sequencing yields millions of sequences, or metabarcodes, that represent the target gene region for the entire sample. The resulting sequences can be identified by matching to a database of sequences for known zooplankton species.
Using metabarcoding to analyze zooplankton diversity routinely and reliably poses many challenges. However, a growing community of researchers is working to build reference databases that link DNA barcodes to species names for zooplankton collected anywhere and everywhere throughout the global ocean.
A global effort
Scientists expect that DNA barcoding and metabarcoding will significantly revise global estimates of zooplankton diversity. But we can only realize the remarkable promise of these approaches through global-scale conversation, cooperation and collaboration.
Several international collaborative efforts have paved the way. They include the 2004-2010 Census of Marine Zooplankton, which identified thousands of species and described hundreds of new species previously unknown to science. I am a member of the Working Group on Integrated Morphological and Molecular Taxonomy, which is carrying this work forward. Deeper insights into zooplankton biodiversity will provide a foundation for future research, monitoring and management of the largest habitat on Earth – the open ocean.
Provided by The Conversation
This article was originally published on The Conversation. Read the original article.![]()